Protein purification and analysis using column chromatography is facing new challenges
because of increasing demands in the proteomic
and biotechnology fields. Column chromatography
involves binding and elution of proteins
with stationary phase, whether such binding is
intentional or accidental. Tight binding may
cause problems in the elution of proteins. Nonspecific
protein binding causes protein loss
and damage to the columns. The authors have
observed that the addition of arginine to column
solvents (mobile phase) improves recovery
and separation of proteins by reducing interaction
of the protein with the column.1–3 In addition,
arginine prevents protein molecules from
interacting with themselves or other molecules
and hence reduces aggregation.4–7 In this paper,
the effects of arginine on the performance of
Protein A, antigen-affinity, hydrophobic interaction, ion-exchange, and size exclusion chromatographies
will be reviewed.
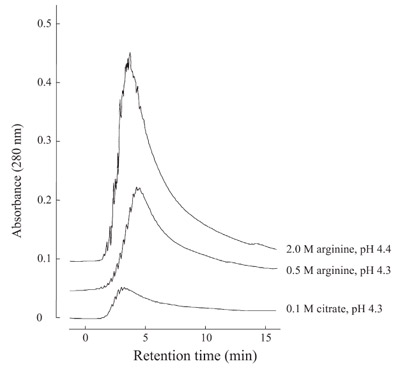
Figure 1 - Protein A column chromatography of monoclonal antibody. Purified
MAb was loaded onto a 1-mL HiTrap Protein A column (GE Healthcare,
Tokyo, Japan) at neutral pH and eluted with the solvents indicated.
Protein A binds to the Fc region of antibodies
and is therefore used to capture antibodies and
recombinant proteins fused with Fc (Fc-fusion
proteins). Both antibodies and Fc-fusion proteins
are versatile reagents for the analysis of expression
and function of physiologically important
proteins. They are also developed as pharmaceutical
drugs. Fc binds to Protein A columns
so tightly that elution of the proteins requires
a harsh condition (e.g., low pH). In order to
circumvent this shortcoming, various
low-affinity Protein A mimics have
been developed.8,9 These, however, suffer
greatly compromised purification
efficiency. The authors have observed
that arginine used as a low-pH solvent
enhances the elution of antibodies from
Protein A.1,2 Figure 1 shows a comparison
of a conventional citrate buffer with
aqueous arginine solution upon elution
of a monoclonal antibody. Since the
antibodies and the structure of the Fc
domain undergo conformational changes
and have reduced stability as the elution
pH is lowered, it was attempted to elute
the bound antibodies at or above pH
4.0. There is little elution observed with
0.1 M citrate at pH 4.3, while both 0.5
and 2.0 M arginine resulted in greatly
enhanced elution.
Antigen-affinity
chromatography
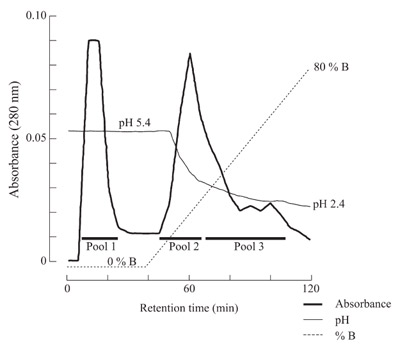
Figure 2 - Antigen-affinity chromatography of antisera against interleukin-
6. Interleukin-6 was conjugated to a 1-mL NHS-HiTrap column, to which
antisera against interleukin-6 was loaded. The bound proteins were eluted first
with 2 M arginine at pH 5.4 (A) followed by a linear gradient of 100% A to
100% B (2 M arginine at pH 2.4).
Polyclonal antibodies can be purified
by antigen-affinity chromatography.
Depending on the affinity of the antibodies
for the antigen, a harsh elution condition such
as low pH may be required. In fact, such high-affinity
antibodies, which are hence difficult to
elute, may be the better reagents. Although the
authors have not done a comparison of arginine
elution with conventional elution, they have
shown that an acidic aqueous solution of arginine
can be used to elute antibodies bound to
the antigen columns. Figure 2 shows the
elution of polyclonal antibodies from
an antigen-conjugated agarose column
using arginine. In this experiment, antisera
raised against interleukin-6 were
bound to the antigen column and the
bound antibodies eluted with a descending
pH from 5.4 to 2.4 in 2 M arginine.
Multiple elution peaks are observed,
presumably due to different affinity
for the antigen; the pooled antibodies
(Pool 1–3) all showed that they bind
interleukin-6 and consist of heavy and
light chains.
Hydrophobic interaction
chromatography
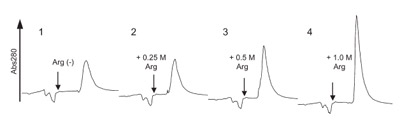
Figure 3 - HIC of interleukin-6. Purified interleukin-6 in 1 M ammonium
sulfate containing 0.5 M arginine was loaded onto a 1-mL HiTrap phenyl-sepharose
column (high phenyl density) in the same solvent and eluted with
0.25 M ammonium sulfate in the presence of 0, 0.25, 0.5, and 1.0 M arginine
(indicated by an arrow).
Hydrophobic interaction chromatography (HIC) uses weakly hydrophobic
ligand to capture the proteins in the
native state through hydrophobic interaction.
Because of weak hydrophobicity of
both the column and the native protein,
most proteins require ammonium sulfate for binding to HIC columns, such as phenyl-sepharose.
The bound proteins are eluted by lower
concentrations of ammonium sulfate, but with
potential loss and aggregation of the proteins.
The authors have tested the ability of arginine to
enhance elution. Inclusion of 0.5–1 M arginine in
the loading samples resulted in weaker binding of
recombinant interleukin-6 (rhIL-6) and a monoclonal
antibody using low-substituted phenyl-sepharose,
causing a portion of the proteins to flow
through the column at 2 M ammonium sulfate,
at which binding was complete in the absence of
arginine.10,11 It is evident that arginine reduces
binding of proteins to the hydrophobic ligands.
Arginine at 0.5 M had no effect on binding when
high phenyl density phenyl-sepharose was used.
However, arginine did facilitate the elution of
interleukin-6 from high phenyl density phenyl-sepharose.
Thus, interleukin-6 was loaded onto
phenyl-sepharose in 1 M ammonium sulfate containing
0.5 M arginine and eluted with a step elution
of 0.25 M ammonium sulfate. Inclusion of
0.25–1 M arginine into the 0.25 M ammonium
sulfate resulted in a sharper elution peak, as shown
in Figure 3. Since the recovery was already close to
100% using ammonium sulfate alone at 0.25 M or
less, it did not significantly increase with the addition
of arginine.
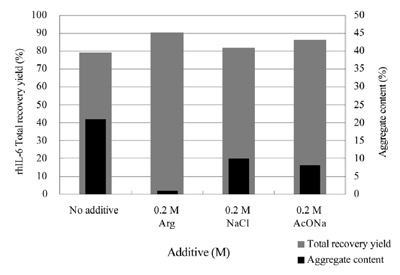
Figure 4 - IEC of interleukin-6. Forty milligrams of refolded and semipurified
interleukin-6 supplemented with 0.2 M of each salt was loaded onto a 20-mL CM-Sepharose FF column (GE Healthcare), 1.6 × 10 cm, and eluted
by the linear gradient of sodium acetate concentration. Detailed conditions are
described in a previous report.10
Arginine is ionic and must be used with caution in ion-exchange chromatography (IEC). Nevertheless,the advantage of including arginine in the loading
sample was obvious in a few applications. Following
are the results seen with interleukin-6.10,11 Refolded
interleukin-6 was loaded in the absence and presence
of arginine (and other salts for comparison), whose
concentration was far below the ionic strength used to
elute the protein from the same column. The bound
protein was eluted without arginine by raising the ionic
strength. Figure 4 shows a comparison of recovery and
aggregate content of eluted materials with and without
arginine included in the loading sample. Recovery
without arginine or other salts was about 80%, but the
aggregate content was high, above 20%. The recovery
increased slightly and the aggregate content decreased
significantly to 5–10% when the loading sample contained
0.2 M NaCl or sodium acetate (AcONa). Inclusion
of 0.2 M arginine further improved both recovery
(to 90%) and aggregate content. In particular,
aggregate decreased to nearly zero. Since arginine is
incapable of dissociating such aggregates, the loading
sample probably did not contain aggregates. Aggregates
were generated during sample loading or elution,
when arginine or other salts are not included. Salts presumably
caused an effect via their ionic strength—i.e.,
they weakened ionic interactions between the proteins
and ion-exchange column, thereby reducing overconcentration
of the loaded protein at the top of the
column (since protein samples are pumped into the
column). Arginine, in addition to contributing to ionic
strength, as is the case for NaCl and AcONa, served
to reduce both protein–protein and protein–stationary
phase interactions.
Size exclusion chromatography (SEC) is a
workhorse for characterization of pharmaceutical proteins. It is the standard
technique for determining the
amount of aggregated forms in
the final product of pharmaceutical
proteins. It is increasingly
critical to have the ability
to accurately assess the amount
of aggregates in pharmaceutical
products, since aggregates
cause various toxic side effects.
For example, immunoglobulin
aggregates have long been
known to cause anaphylactoid
reactions. More recently, a serious
concern regarding aggregation
was raised after an upsurge
in the incidence of pure red
cell aplasia (PRCA), although
there is no conclusive evidence
of aggregation of erythropoietin
(or formation of aberrant
structure) being involved in this
incidence. While simple and
high in throughput, SEC has a
problem with nonspecific binding,
i.e., the proteins, in particular
aggregated proteins, bind to the
SEC columns nonspecifically, leading
to an incorrect estimate of aggregation,
insufficient separation of protein species,
or coelution of proteins with low
molecular weight solvent components.
These problems can be overcome by
adding arginine to the elution (mobile
phase) solvent.
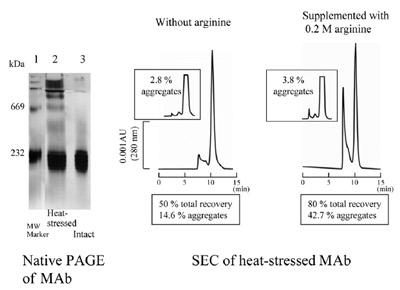
Figure 5 - SEC of monoclonal antibody. Aggregated mouse MAb (4.66 μg
measured by UV absorbance at 280 nm) was applied to a TSK G3000SWXL
column (Tosoh, Tokyo, Japan) equilibrated with 0.1 M sodium phosphate at
pH 6.8 with or without 0.2 M arginine hydrochloride. Intact MAb (4.66 μg)
was analyzed in each condition (inset). Aggregated MAb sample was generated
by heating the purified MAb preparation. Native gel of aggregated MAb (2 μg)
or intact MAb (1 μg) was carried out on a Phastsystem using Phastgel 7.5%T
(GE Healthcare).
Figure 5 shows an example using a monoclonal
antibody. Native gel analysis shows that the
unstressed sample (lane 3) is homogeneous and
the stressed sample (lane 2) contains a large
amount of aggregates. When these samples were
subjected to SEC analysis in the absence of arginine,
the recovery of the eluted proteins was
only 50%. In addition, the aggregate content is
only 14.6%, well below the amount indicated by
visual inspection of the native gel data. When
0.2 M arginine was included, the recovery of
eluted proteins jumped to 80% and the aggregate
content to 42.7%. Thus, arginine resulted in
enhanced elution of both monomeric and aggregated
forms of the antibody. The same was true
for the unstressed antibody sample (inset in Figure
5). In the absence of arginine, the aggregate
content was 2.8%, while in 0.2 M arginine, it was
3.8%; in particular, 0.2 M arginine resulted in
enhanced elution of larger aggregates, shown in
the bottom-right panel of Figure 5.
Is arginine safe?
There are of course other reagents that can be
used for the above purpose. One reason why
arginine was chosen is its inertness to proteins.
Arginine does not denature proteins at or below
room temperature. It also does not disaggregate
or dissociate stable complexes, as shown
in its inability to dissociate antibody–antigen
or antibody–Protein A complexes. It has been
observed that the aggregates separated by SEC
in the presence of arginine do not dissociate.
In addition, bovine serum albumin, which does
not bind to the column, gave identical amounts
of oligomers in the absence and presence
of arginine.
References
- Arakawa, T.; Philo, J.S.; Tsumoto, K.; Yumioka, R.;
Ejima, D. Protein Expr. Purif.2004, 36, 244–8.
- Ejima, D.; Yumioka, R.; Tsumoto, K.; Arakawa, T.
Anal. Biochem. 2005, 345, 250–7.
- Ejima, D.; Yumioka, R.; Arakawa, T.; Tsumoto, K. J.
Chromatogr. A 2005, 1094, 49–55.
- Arakawa, T.; Tsumoto, K. Biochem. Biophys. Res.
Commun. 2003, 304, 148–52.
- Shiraki, K.; Kudou, M.; Fujiwara, S.; Imanaka, T.;
Takagi, M. J.Biochem. 2002, 132, 591–5.
- Umetsu, M.; Tsumoto, K.; Nitta, S.; Adschiri, T.;
Ejima, D.; Arakawa, T.; Kumagai, I. Biochem. Biophys.
Res. Commun. 2005, 328, 189–97.
- Tsumoto, K.; Umetsu, M.; Kumagai, I.; Ejima, D.;
Philo, J.S.; Arakawa, T. Biotechnol. Prog. 2004, 20,
1301–8.
- Li, R.; Dowd, V.; Stewart, D.J.; Burton, S.J.; Lowe,
C.R. Nat. Biotechnol. 1998, 16, 190–5.
- Fassina, G.; Verdoliva, A.; Palombo, G.; Ruvo, M.;
Cassani, G. J.Mol. Recognit. 1998, 11, 128–33.
- Ejima, D.; Watanabe, M.; Sato, Y.; Date, M.; Yamada,
N.; Takahara, Y. Biotechnol. Bioeng.1999, 62, 301–10.
- Ejima, D.; Arakawa, T.; Tsumoto, K., in preparation.
Mr. Ejima is in the Applied Research Dept., AminoScience
Laboratories, Ajinomoto Co., Inc., Kawasaki, Japan. Dr.
Tsumoto is in the Dept. of Medical Genome Sciences, Graduate
School of Frontier Sciences, The University of Tokyo,
Kashiwa 277-8562, Japan; e-mail: [email protected].
Dr. Arakawa is with Alliance Protein Laboratories, Thousand
Oaks, CA, U.S.A.