Recent developments in upstream processing of
biomolecules, driven by an increasing demand for
biotherapeutics, have resulted in increased protein
expression levels and larger feed volumes. As the
total amount of protein increases, the capture and
intermediate purification steps in the downstream
process must be able to handle larger volumes with
higher amounts of expressed protein in a fast and
efficient way. This requires separation media with
properties supporting both high flow velocity and
high dynamic binding capacity.
The Capto™ Q is a strong anion exchanger (GE
Healthcare Bio-Sciences AB, Uppsala, Sweden) based
on a highly rigid agarose matrix. It combines high
capacity with high flow rates at low backpressures.
This article describes the capture of green fluorescent
protein (GFP) on Capto Q. Results from
media screening, optimization, scaleup, as well as
scale-up modeling and productivity calculations
are presented. In addition, some general guidelines
on optimizing a capture step when using the anion exchanger are given.
GFP is a 28-kDa protein with a pI of 6.2, in nature
found in the pacific jellyfish AequoreaVictoria. In
this study, recombinant GFP expressed in Escherichiacoli was purified. The goal of the study was to demonstrate
that it is possible to capture 100 kg of product
per 24 hr using Capto Q. As will be shown, the goal
was reached.
Experimental and results
Method development and optimization work was
performed using an ÄKTAexplorer™ system, while
the scaleup was performed using an ÄKTApilot™
system (both from GE Healthcare Bio-Sciences
AB). Since GFP absorbs light specifically at 490
nm, it was easy to follow the protein during process
development and optimization.
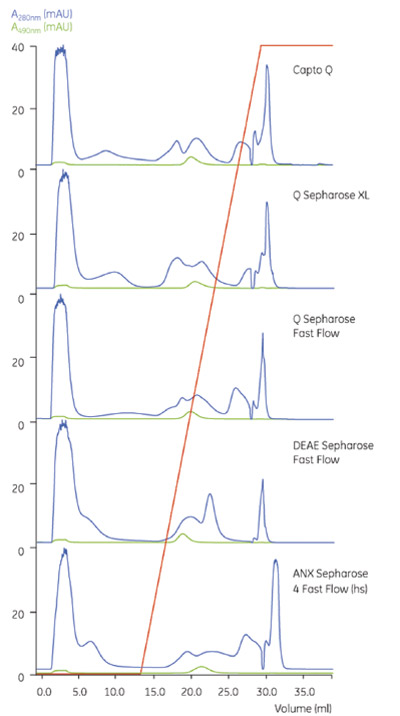
Figure 1 - Screening of media selectivity. From top to bottom:
Capto Q, Q Sepharose XL, Q Sepharose Fast Flow,
DEAE Sepharose Fast Flow, and ANX Sepharose 4 Fast Flow
(hs). Blue curve: A280, green curve: A490 (GFP). (Figures
1–4 © 2006 GE Healthcare Bio-Sciences AB. Reproduced
with kind permission.)
An initial benchmarking of different
anion exchange media (Capto Q, Q
Sepharose™ Fast Flow, Q Sepharose
XL, ANX Sepharose 4 Fast Flow [hs],
and DEAE Sepharose Fast Flow [all
from GE Healthcare Bio-Sciences
AB]) was done at small scale using
prepacked HiTrap™ columns (GE
Healthcare Bio-Sciences AB). Figure
1 shows the result of the screening.
It can be seen that Capto Q gave a
selectivity profile that differed slightly
from the other tested media and it was
decided to use Capto Q for optimization
of the capture step.
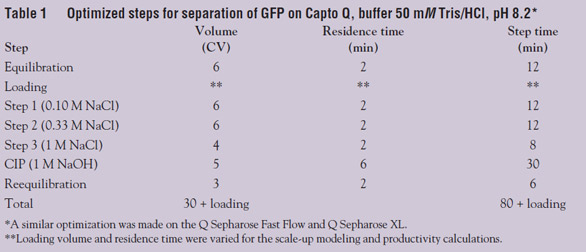

Optimization of the capture step was
performed, determining both optimal pH and conductivity for binding and elution as
well as residence time for optimal productivity. To
save process time and minimize buffer consumption, the process equilibration/reequilibration/wash/elution/cleaning in place (CIP) volumes
were optimized without sacrificing purity and yield.
It was found that the optimal loading buffer was 50
mM Tris/HCl, pH 8.2. Elution was accomplished
by using a step gradient with an increasing amount
of NaCl. Table 1 shows the optimized steps for the
separation. One should be aware that for some
proteins, the dynamic binding capacity on Capto
Q does increase upon increased ionic strength up
to a certain ionic strength, over which it again
decreases. It is therefore recommended that both
loading pH and loading conductivity, between
2 and 15 mS/cm, are optimized with respect to
dynamic binding capacity as well as purity of the
target protein.
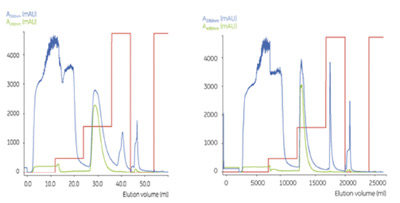
Figure 2 - Capture of GFP on small-scale (2-mL) and pilot-scale (800-mL)
columns. On the small scale, 30 mg, and on the pilot scale, 12 g of GFP were
loaded per run. Note that in the pilot-scale chromatogram, the volume axis includes
the equilibrium step; in the small-scale experiment, the start point is at the start of
sample introduction.
The developed method was used for scaling up
the separation to pilot scale. A 400-fold scaleup
was performed from a Tricorn™ 5/100 column
(GE Healthcare
Bio-Sciences AB) (10-cm bed
height, 2-mL volume) to a FineLINE™ 70 column
(GE Healthcare Bio-Sciences AB) (20-cm bed
height, 800-mL volume), keeping the residence
time constant. Figure 2 shows a comparison of the
results from the two scales. The purification factor
was at both scales approx. 5, and yields in all runs
were above 90%. Analysis by 2-D electrophoresis
showed that the profile of the copurified contaminants
was very similar at the two scales (data not
shown). Scaleup of processes for biopharmaceuticals
requires that the separation profile remain the
same at the different scales. This includes parameters
such as purification factor and yield as well as
clearance of critical impurities like nucleic acids,
host cell proteins, and viruses. Any of these factors
may change when column bed height is altered,
and should thus be validated using the final bed
height. This is of great importance to remember,
since in the early stages of a project there is often
a shortage of material and work is carried out in
columns with lower bed heights than will be used
at the production scale.
Scale-up calculations
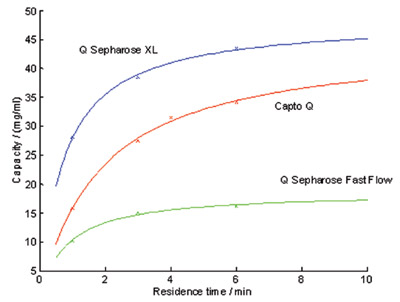
Figure 3 - Dynamic binding capacities for GFP on Capto Q, Q Sepharose XL,
and Q Sepharose Fast Flow as a function of residence time.
Dynamic binding capacities (DBC) for GFP
were determined at different residence times
on Capto Q, Q Sepharose XL, and Q Sepharose
Fast Flow (Figure 3). Based on these results and
the chromatography cycle data of
the three media (parameters used for
Capto Q shown in Table 1), productivity
calculations for each medium
were performed.
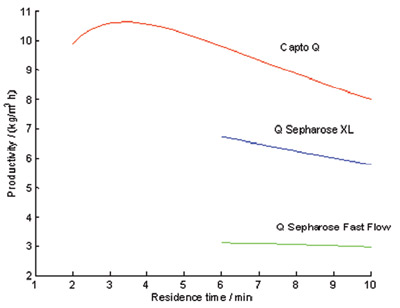
Figure 4 - Calculated productivity with Capto Q compared to Q Sepharose XL
and Q Sepharose Fast Flow. Note that Q Sepharose XL and Q Sepharose Fast
Flow do not have the same rigidity as Capto Q; thus there are no values for these
media at the shorter residence times attainable at the 20-cm bed height.
A theoretical scaleup was done to 20-cm bed height by keeping the respective
residence times constant, i.e., operating
Capto and Sepharose (Fast Flow and
XL) resins at 600 cm/hr and 200 cm/hr,
respectively. Exceptions were the loading
step, run at different flow velocities,
and the CIP, which was kept constant
at 30 min. The cycle time when using
Capto Q (1.5 hr) is considerably shorter
compared to when using Q Sepharose
XL or Q Sepharose Fast Flow (3.5 hr).
The columns were loaded to 70% of
DBC (QB 10%) and yield was measured
to be 93.5%. The effect of varying the
loading velocity on resin productivity is
shown in Figure 4. Since Capto Q allows
higher flow velocities to be used, the
developed process can be run at shorter
residence times compared to Q Sepharose
XL or Q Sepharose Fast Flow. Thus,
note that although the DBC is higher
on Q Sepharose XL compared to Capto
Q (Figure 3), the productivity is higher
on Capto Q. The high productivity
achieved on Capto Q stems from both
a high dynamic binding capacity and
the possibility of using high flow velocities,
even at a large scale. As can be seen
from Figure 4, the maximum productivity
achieved on Capto Q is 10.4 kg/m3hr.
This corresponds to the goal of 100 kg of
product per 24 hr in a 400-L column (0.2
× 1.6 [i.d.] m).
Conclusion
A capture step was developed that
gave a high yield of target protein as
well as removal of bulk impurities in less than
2 hr. A 400-fold scaleup was demonstrated.
Recoveries were above 90% at both scales. The
purification factor, approx. 5, indicates that the
majority of contaminants had been removed.
Not only were the recoveries high, but the product
eluted in a small volume (2.6 column volumes
in the FineLINE 70 column), which is
critical for a subsequent chromatography step.
Finally, a short overall process time means
improved product integrity by reducing exposure
time to proteases and other contaminants detrimental
to product quality and yield. Altogether,
the results show that Capto Q is well suited for
use in a capture step.
Ms. Åkerblom and Mr. Bryntesson are Research Engineers,
Dr. Brekkan is a Scientist, and Dr. Eriksson is a Senior Scientist,
GE Healthcare Bio-Sciences AB, Björkgatan 30, SE-
751 84 Uppsala, Sweden; tel.: +46 18 6120539; fax: +46
186121844; e-mail: [email protected].