Over the years, researchers have used density gradient
techniques for separating viruses and macromolecular
species such as proteins, subcellular organelles, and
nucleic acids. Brakke first used these techniques,1 and
a publication by Griffith2 provides a thorough discussion
of the rate zonal and isopycnic density gradient
procedures. Both fixed angle and swinging bucket
rotors have been used for these two methods. The
vertical tube rotors with the tube angle fixed parallel
to the axis of rotation are also being used for density
gradient studies in the ultracentrifuge.
This paper compares separations of similar samples
obtained with these four rotor designs. Run times for
the rate zonal method were calculated to provide the
best resolution for each rotor. Additionally, the rotor
instruction manuals were used to obtain optimum
rotor speeds for isopycnic banding of DNA in cesium
chloride (CsCl) gradients.
Experimental
The models L5-75 and L8-80 ultracentrifuges
(Beckman
Coulter, Fullerton, CA) were used.
Rotors used were the SW 55Ti and SW 28 swinging
bucket rotors (Beckman Coulter); F65L-6 × 13.5 mL
and F50L-8 × 39 mL fixed angle rotors (FIBERLite
Centrifuge Inc., Santa Clara, CA); and VTi 65,
NVT 65, and VTi 50 vertical tube rotors (Beckman
Coulter). After centrifugation, the gradients were
monitored as discussed by Griffith.2
Rate zonal separation (sucrose gradients)
-
Proteins: a) Bovine serum albumin (BSA), fibrinogen,
and a mixture of BSA and fibrinogen were placed on
sucrose gradients (10–40% w/w) in the F65L-6 × 13.5
mL, VTi 65, and SW 55Ti rotors. The rotors were run
at maximum speeds for 3 hr, 3 hr, and 16 hr, respectively.
The run temperature was 5 °C. The short-column
(half-filled or short tubes) method by Griffith2
was used in the F65L-6 × 13.5 mL rotor, while
the VTi 65 and SW 55Ti rotors had completely filled
tubes. Sample volumes were the same in each rotor
(0.2 mL, 1%). b) The two molecular forms of human
mammary estrogen receptors (4S and 8S) were run in
the F65L-6 × 13.5 mL and NVT 65 rotors. Sucrose
gradients (10–40% w/w) were used and the rotors had
similar loads (0.2 mL). Maximum rotor speeds were
maintained for 3 hr and 2.5 hr, respectively. The run
temperature was 5 °C.
A separate experiment was done to show the shifting of
the 8S molecular form of the receptor to the 4S position
when 0.4 M KCl was added to the sample and gradient.
After centrifugation, the model 250 liquid scintillation
counter (Beckman Coulter) was used to identify the
separated components. No studies were done with
the swinging bucket rotor because this separation is
documented in the literature. Run times for similar
gradients in swinging bucket rotors with full tubes
would have been 16 hr at 5 °C. Pavlik et al.3–5 used
short-column methods for estrogen receptors, and
run times for these studies were 3 hr.
- Subcellular particles (polysomes from rat liver):
Rat liver polysomes prepared by the method of
Noll6 were run on a 10–40% w/w sucrose gradient
in the VTi 50, F50L-8 × 39 mL, and SW 28
rotors. The short-column method and similar
tubes were used in the F50L-8 × 39 mL and SW
28 rotors. The VTi 50 rotor had full tubes. The
sample load was placed on the gradients in the
three rotors (2 mL, 40 μg/mL). At 5 °C, rotor
speeds and run times were as follows: SW 28
rotor—27,000 rpm for 150 min, F50L-8 × 39
mL rotor—50,000 rpm for 40 min, and VTi 50
rotor—50,000 rpm for 35 min.
Isopycnic banding of DNA
- Two DNAs—Micrococcus luteus (1.73 g/
mL) and λ bacteriophage (1.71 g/mL)—were
mixed in tris–EDTA (ethylenediaminetetraacetic
acid) buffer to obtain an absorbance of
2.0 AUFS at 260 nm. The rotors chosen to
observe the separation of this mixture were
the SW 55Ti, F65L-6 × 13.5 mL, and NVT
65. The density of the buffered DNA mixture
was adjusted with CsCl to 1.60, 1.65, and 1.70
g/mL, respectively, and the run times for the
rotors were 16 hr. In the NVT 65 and F65L-6 × 13.5 mL rotors, each tube was loaded with 13.5
mL of CsCl solution, whereas 2 mL was loaded in the
SW 55Ti rotors and run at 50,000 rpm. The NVT 65
and F65L-6 × 13.5 mL rotors were run at 65,000 rpm,
and the temperature for each run was 20 °C.
The DNA experiment in the NVT 65 and F65L-6 ×
13.5 mL rotors was repeated to observe separations in
5 hr. The F65L-6 × 13.5 mL rotor was loaded with 5.0
mL CsCl solution to use the short-column method.
- Samples of crude lysate from E. coli bacteria containing
plasmids pBR 322 were prepared by the method of
Katz et al.7 CsCl solutions were adjusted to densities
of 1.55 g/mL, and 3 mL of the nucleic acids containing
ethidium bromide (EtBr) solution were added to
the CsCl density to fluoresce the DNA. The solutions
were run in the VTi 65 and NVT 65 rotors at their
maximum speeds, and the run time and temperature
were 16 hr at 20 °C; the F65L-6 × 13.5 mL was run at
60,000 rpm at the same temperature with filled tubes.
After the centrifugation was terminated, long-wave
UV light was used to fluoresce the DNA zones in
the centrifuge tubes. The lower zone containing the
plasmid DNA was removed by piercing the tube walls
with a hypodermic needle and syringe. Thin-walled
polyallomer tubes manufactured by Seton Scientific
Co. (Los Gatos, CA) were used for this purpose. The
EtBr was extracted from the plasmids with CsCl saturated
n-butanol followed by low-speed centrifugation
(6000 rpm for 15 min). The recovered pellet was
resuspended in tris–HCl buffer and analyzed for purity
by agarose gel electrophoresis.
Results and discussion
Rate zonal separations (sucrose gradients)
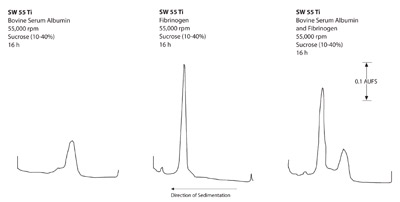
Figure 1 - Rate zonal separations of BSA and fibrinogen in the SW
55Ti swinging bucket rotor.
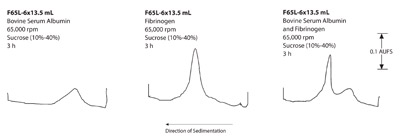
Figure 2 - Rate zonal separations of BSA and fibrinogen in the
F65L-6 × 13.5 mL fixed angle rotor.
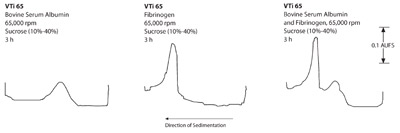
Figure 3 - Rate zonal separations of BSA and fibrinogen in the VTi
65 rotor.
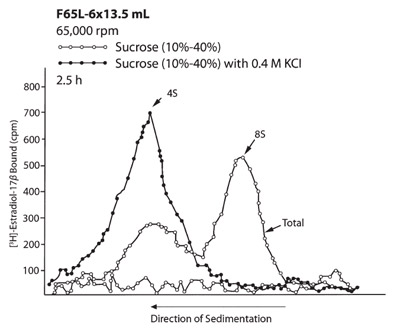
Figure 4 - Rate zonal separations of the two molecular
forms of estrogen receptors from human mammary tumor in
the F65L-6 × 13.5 mL fixed angle rotor.
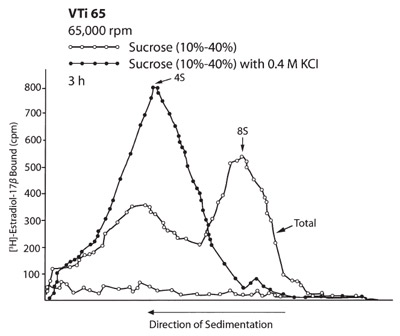
Figure 5 - Rate zonal separations of the two molecular
forms of estrogen receptors from human mammary tumor in
the VTi 65 rotor.
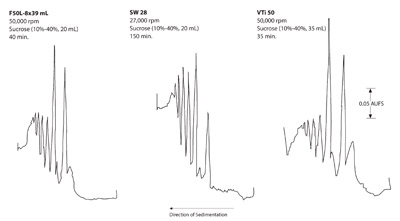
Figure 6 - Rate zonal separation of polysomes from rat liver.
- Proteins: a) Figures 1–3 show that the swinging
bucket rotor provided the best separation of the
three rotors, but with the longest run times. Shorter
run times are possible with the short-column
method
without a reduction in sample load. Studies by Pavlik
et al. have shown that the run times for proteins
in swinging bucket rotors have been reduced by 75%
when the short column method is used.3 b) Figures 4
and 5 show that both run times were similar in the
VTi 65 and F65L-6 × 13.5 mL rotors. However, the
separated zones in the vertical tube rotor were much
wider than in the F65L-6 × 13.5 mL rotor because
the zones were separated in larger gradient volumes
than in the fixed angle rotor. This corresponds to
the geometric differences in the gradient reorientation
process between the F65L-6 × 13.5 mL rotor
and NVT 65 rotor. The result is therefore better
separation of the zones with the fixed angle rotor.
- Subcellular particles (polysomes): Figure 6 shows the
separation of polysomes in the three rotors. Again,
the swinging bucket rotor provided the best separation
of the three. Although the same sample load was
used in the three rotors, the peak heights of the polysomes
were taller in the swinging bucket rotor than in
the F65L-6 × 13.5 mL rotor. However, there was loss of the polysomes in the vertical tube after the zones
moved toward the centrifugal area during separation.
Isopycnic banding (DNA
samples)
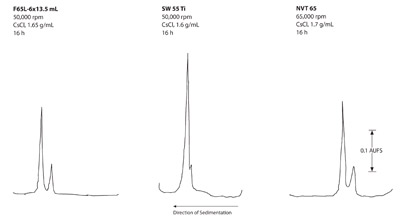
Figure 7 - Isopycnic separations of DNA from Micrococcus
luteus and λ bacteriophage.
Figure 7 shows the separations of the two DNAs in the
three rotor types. Comparable separations in the same
run times were observed in the near vertical tube and
fixed angle rotors. The swinging bucket rotor did not
completely separate the two components.
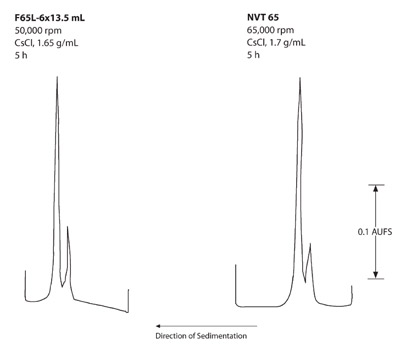
Figure 8 - Isopycnic separations of DNA from Micrococcus
luteus and λ bacteriophage.
The 5-hr runs in Figure 8 show less separation of the
DNA than the 16-hr runs in Figure 7; however, this
separation may be acceptable for some experiments.
During tube fractionation, the volume of CsCl solution
separating the two DNA zones in both rotors for the
16-hr run was measured. The zones in the NVT 65 rotor
had a slightly larger volume (0.5 mL) than the F65L-6 ×
13.5 mL rotor (0.4 mL). This corresponds to the geometric
differences between the reorientation processes in the
two rotors. Additionally, the reorientation process is less
in the F65L-6 × 13.5 mL rotor than in the NVT 65, and
the peak-to-valley ratio is better in the fixed angle rotor
than in the near vertical tube rotor. This resulted in better
separation of the zones with the fixed angle rotor.

Figure 9 - Purification of pBR 322 plasmids from E. coli by buoyant density gradient centrifugation. Note the RNA contaminant
falling off the centrifugal tube wall after the run in the vertical tube (a) and near vertical tube (b) rotors. The contaminant is pelleted in
the fixed angle (c) rotor during the run (see text).
Photographs of the fluorescent EtBr–DNA zones separated
in the vertical tube, near vertical tube, and fixed
angle rotors are shown in Figure 9a–c. The upper zone
represents chromosomal DNA, while the lower zone represents
plasmid DNA. EtBr also binds to the RNA from
the crude lysate. The RNA contaminant is observed pelleted
at the bottom of the tube in the F65L-6 ×13.5 mL
rotor in Figure 9c. The flocculent contaminant RNA is
observed falling from the centrifugal wall of the vertical
tube rotor VTi 65 in Figure 9b, and the RNA contaminant
is observed falling from the wall of the near vertical
tube rotor NVT 65 in Figure 9a.
Conclusion
Rate zonal separation (sucrose gradients)
When proteins are run in sucrose gradients, the swinging
bucket rotors provide slightly better separation
between components. Since these rotors have the
longest pathlength, the run times will be longer than
in the vertical tube and fixed angle rotors. During rate
zonal separations, particles are separated according to
size differences. With multicomponent systems (more
than two sample components) or when the sedimentation
coefficient differences are very small (less than
4S units), the rotor with the longest pathlength and
the highest centrifugal force should be chosen for the
best separation between components. If shorter run
times are essential, the short-column method2 can be
used in swinging bucket rotors. The fixed angle rotor
will also provide short run times; however, the separation
between components will be better than in the
near vertical or vertical tube rotors.
The swinging bucket rotors are therefore recommended
for sucrose gradients when the best separation is needed.
If shorter run times are essential, the short-column
method2 can be used in swinging bucket rotors. The
fixed angle will also provide short run times; however,
the separation between components will be better than
in the near vertical or vertical tube rotors.
Isopycnic banding (cesium
chloride gradients)
In this method, samples such as nucleic acids separated
according to their density differences in cesium chloride
solutions. After the zones have reached their isopycnic
position in the tubes, there is no further sedimentation
through the gradient. At this point, the zones cease to
spread. The best separation therefore depends on the
volume of solution that separates the sample zones at the
end of the run. The geometry of the rotors is such that
during centrifugation, the vertical tube or near vertical
tube rotor has the largest volume of solution separating
two zones. Flamm et al.8 and Fisher et al.9 demonstrated
that swinging bucket rotors have the smallest volume of
solution separating the two zones since there is no reorientation
of the solution during deceleration of the rotor.
The fixed angle rotor gave similar separations in the
same run time as the vertical tube rotor; however,
there is a greater than 0.1-mL difference in the volume
of CsCl solution separating the two DNA zones
as shown in the NVT 65 and the F65L-6 × 13.5 mL
rotors. Gradient reorientation in the NVT 65 rotor
caused the zones to be larger, thus giving a loss of
resolution, as shown in Figures 7 and 8.
When the fixed angle and vertical tube rotors were used
to separate plasmid DNA, the run times were similar.
With the fixed angle rotor, the contaminating RNA was
pelleted at the bottom of the tubes and away from the
purified plasmid DNA in a single run. This was reported
by Wong et al.10 and Griffith.11 Little12 reported that the
RNA contaminant fell off the tube wall of the NVT 65
rotor during centrifugation. The author also mentioned
that varying the concentrations of Triton X-100 (Rohm
and Haas, Philadelphia, PA) in the sample preparation
could prevent the RNA contaminant from adhering to
the centrifuge tube walls. However, using Triton X-100
to prevent RNA from adhering to the tube wall would
not prevent the plasmid DNA from being contaminated with the RNA. Maniatis et al.13 reported that minor
quantities of contaminating RNA greatly reduce the
specific activity of 32P-end labeled DNA required for
Maxam-Gilbert DNA sequencing.14 This was verified by agarose gel electrophoresis.
Investigators have been using a modified prepurification
procedure for plasmid isolation. The procedure
adds a phenol extraction followed by ethanol precipitation
of the DNA to the standard polyethylene
glycol method reported by Katz et al.7 Since all of the
contaminating protein and RNA are not removed
by the extraction, the residual contaminants often
recontaminate the sample by falling from the wall of
the tube before the plasmid band can be extracted. For
this reason, users of vertical tube rotors may perform
a second overnight run to further purify the plasmid
DNA. The fixed angle rotors are therefore recommended
for plasmid DNA separations, especially when
it is essential to remove all contaminants. The higher
force fields generated by most fixed angle rotors permit
separations to be completed in run times equal to
those made in the vertical or near vertical tube rotors.
Reduced run times at the higher g-forces for fixed
angle rotors can be calculated for short pathlength
(cone top) polyallomer tubes using the K-Factor formula
for these tubes in the fixed angle rotors. The
tubes are manufactured by Seton Scientific Co.
References
- Brakke, M.K. Density gradient centrifugation: a new
separation technique. J. Am. Chem. Soc.1951, 73,
1847–8.
- Griffith, O.M. Techniques of Preparative, Zonal, and
Continuous Flow Ultracentrifugation, 3rd ed. Beckman
Instruments, Inc., Spinco Div.: Palo Alto, CA, 1979.
- Pavlik, E.J.; Rutlege, S. Estrogen-binding properties of
cytoplasmic and nuclear estrogen receptors in the presence
of Triton X-100. J. Steroid Biochem. 1980, 13, 1433–41.
- Griffith, O.M. Rapid density gradient centrifugation
using short column techniques. Anal. Biochem. 1978,
90, 435–43.
- Goral, J.E.; Wittliff, J.L. Comparison of glucocorticoid
binding proteins in normal and neoplastic
mammary tissues of the rat. Biochemistry1975, 14,
2944–52.
- Noll, H. Polysomes: analysis and structure and function.
In Techniques in Protein Biosynthesis, Vol. 2, pp
101–79. Campbell, P.N.; Sargent, J.R., Eds. Academic
Press: New York, NY, 1966.
- Katz, L.; Kingsbury, D.T.; Helinski, D.R. Stimulation
by cyclic adenosine monophosphate of plasmid
deoxyribonucleic acid–protein relaxation complex. J.
Bacteriol. 1973, 114, 577–91.
- Flamm, W.G.; Bond, H.E.; Burr, H.E. Density gradient
centrifugation of DNA in a fixed angle rotor. A higher
order of resolution. Biochem. Biophys.Acta1966, 129,
310–7.
- Fisher, W.D.; Cline, G.B.; Anderson, N.G. Density
gradient centrifugation in angle head rotors. Anal.
Biochem. 1964, 9, 477–82.
- Wong, T.K.; Nicolau, C.; Hofshneider, P.H. Appearance
of β-lactamase activity in animal cells upon lipid
mediated gene transfer. Gene1980, 10, 87–94.
- Griffith, O.M. Rapid isolation of bacterial plasmid
DNA by isopycnic centrifugation in fixed angle rotors.
Applications Data DS-591. Beckman Instruments, Inc.:
Palo Alto, CA, 1981.
- Little, S.E. Plasmid separations in NVTTM near vertical
tube rotors. Applications Data DS-770. Beckman
Instruments, Inc.: Palo Alto, CA, 1998.
- Maniatis, T.; Fritsch, E.F.; Sambrook, J. Molecular
Cloning: A Laboratory Manual. Cold Spring Harbor
Laboratory: Cold Spring Harbor, NY, 1982.
- Maxam, A.M.; Gilbert, W. Proc. Natl. Acad. Sci. USA
1977, 74, 560–4.
Dr. Griffith is Director of Research, FIBERLite Centrifuge Inc.,
422 Aldo Ave., Santa Clara, CA 95954, U.S.A.; tel.: 408-988-1103; 408-988-1196; e-mail: [email protected].