The understanding of protein dynamics,
the way these molecules fold to
form a stable structure, interact with
other molecules, or undergo changes
during enzyme activity, etc., in their
native environment is a key research
area. The general aim is to understand
the underlying fundamental principles
that govern the workings of living
organisms, most of which are carried
out by proteins in interaction with
other molecules. Nuclear magnetic resonance (NMR) spectroscopy, in
particular, is capable of taking proteomics
to the next level, where scientists
study not only the structure,
but also the dynamics and the interactions
of proteins with other molecules.
With NMR, it is possible to gain
insight into how proteins fulfill their
functions in their native aqueous environment.
However, these NMR studies
are challenging and often require
that researchers push the limits by
using the combination of cutting-edge
hardware, such as cold probes, and the
latest innovative new methodology,
such as very fast 2-D NMR.1–5
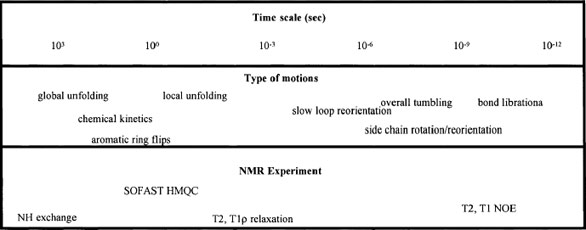
Figure 1 - Protein dynamics—time scale and NMR experiments.
Gathering insight into the dynamic
properties of proteins is challenging.
Very few analytical methods can provide
local structural and dynamic information
for the different protein segments
comprising the protein, such as secondary
structural elements, individual
residues, or even individual nuclei. Further,
besides the need for site-resolved
studies, the time scale of the dynamic
process is also an important factor that
needs to be addressed. The time scale of
the experimental method has to be comparable
to the time scale of the dynamic
process studied (Figure 1). NMR spectroscopy
is uniquely suited to the study of
dynamics of proteins, since it fulfills both
of the above requirements.
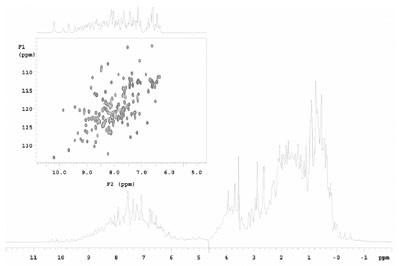
Figure 2 - Comparison of 1-D and 2-D NMR (heteronuclear single quantum coherence [HSQC])
of 3 mM NuiA in 90% H2O 10% D2O.
Figure 1 shows the many types of motions
in proteins that need to be studied and
the matching NMR methods that make
these studies possible. It is clear that
NMR offers a wide range of options to
better understand the dynamic world of
proteins. One significant issue faced by
NMR spectroscopists is that the 1-D
NMR spectrum that can be collected in a
few seconds does not provide the needed
atomic resolution information on large
molecules such as proteins. Researchers
have to turn to 2-D NMR to achieve
site-specific resolution, but the experiment
is too time consuming to meet the
time-scale requirement (Figure 2). The
acquisition time of typical 2-D NMR
spectra ranges from several minutes to
tens of minutes, depending on sample
concentration and the resolution
required in the indirect dimension.
In the past 3–5 years, several exciting fast NMR methods have emerged.1–5 Most of
these methods are intended to accelerate
higher-dimensional (3-D and 4-D) NMR
data collection for protein structure
determination. Unfortunately in most
cases the increase in speed comes at a
price. Namely, shorter experiment times
generally mean that less signal is
acquired. As a consequence, the signal-to-noise ratio (SNR) in fast experiments
in general is dramatically reduced.
Hence, the technology that increases the
SNR is very beneficial. A very fast 2-D
NMR technique, the SOFAST HMQC
(band-selective optimized-flip-angle
short transient heteronuclear multiple
quantum coherence) was introduced in
2005.4,5 The method is distinctive
because it offers a considerable advantage
in speed and potentially increases the
sensitivity per unit time of many conventional
experiments. This allows the
recording of 2-D NMR spectra of proteins
in less than 10 sec, and thus breaks
down the barriers to real-time investigation
of dynamic events in proteins. Several
research groups have recently
demonstrated that the time scale of
hydrogen/deuterium (H/D) exchange
measurements in proteins can be dramatically
increased by combining the sensitivity
advantages of cold probes with fast
NMR methodologies.4–7
SOFAST method
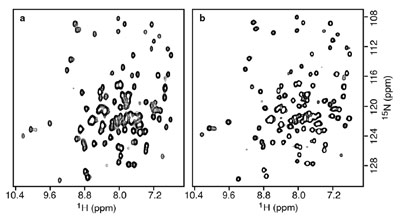
Figure 3 - 2-D 1H-15N correlation spectra of Rb. capsulatus cytochrome C′ (128 residues)
recorded on a Varian 600-MHz NMR spectrometer. a) SOFAST-HMQC with a recycle time of 1
msec and a total experimental time of 6 sec. b) Standard HSQC using a recycle time of 1 sec and a
total experimental time of 180 sec.
In order to reduce the recycling delay that
accounts for most of the experiment time
in conventional multidimensional NMR
spectroscopy, the SOFAST technique
employs both longitudinal relaxation
optimization8 and optimized flip-angle9
band-selective excitation. For instance,the SOFAST-HMQC4,5 experiment
selects the NH (amide) region using a
shaped polychromatic excitation pulse10
that provides a variable flip-angle capability
for the first excitation pulse. This pulse
preserves the +Z magnetization of the
other protein protons (aliphatic and aromatic
sites), which serves as a reservoir of
magnetization to rapidly relax the NH
protons back to thermal equilibrium during
the acquisition time. This setup
makes it possible to set the relaxation
delay to as short as 1 msec versus the traditionally
used relaxation delays of over 1
sec. The reduction in the relaxation delay
from ca. 1 sec to 1 msec decreases experiment
duration by about a factor of 20
(Figure 3), without diminishing the sensitivity
per unit time with respect to conventional
methods.4,5
Unfortunately, most proteins that are
involved in relevant biological processes
are usually available only in very limited
quantities. This problem is further exacerbated
by the fact that protein samples
enriched with at least one magnetically
active isotope (N15 and/or C13) are
required for the basic 2-D correlation
experiments (linking H1 and neighboring
C13 or N15 atoms). The goal of
researchers is to collect data on dilute
protein samples that most closely resemble
the native biological environment,
but in turn they are faced with the
detection limit of their hardware.
The sensitivity of NMR experiments
has always been a challenge. Over the
past 3–5 years, several techniques have
evolved that are helping to push the
barriers of NMR sensitivity and
throughput to new levels. One of the
most exciting and recent developments
in this area is the use of cryogenically
cooled probes.11 These devices have
the electronics within the NMR probe
cooled to approximately –250 °C
(comparable to the temperature on the
surface of Pluto), which results in an
increase in sensitivity by a factor of 2–4
as compared to a conventional NMR
probe. At cryogenic temperatures, the
thermal noise of the radiofrequency
(RF) electronics is significantly
reduced while the probe’s quality factor
is improved. This results in reduced
noise and greater signal, thereby significantly
increasing the overall SNR of
the probe. The improved sensitivity of
the cryogenic probe translates to 4–16
times shorter experiments compared to
data collected on a conventional
probe. This technology was introduced
to the marketplace by Varian (Palo
Alto, CA).12 More importantly, the
company’s cold probe technology has
been developed to achieve the sensitivity
increase without any compromise in
NMR performance.11 As new NMR
methods emerge, however, the robustness
and applicability of the cold probe
becomes challenged.
Combining cold probe technology and
the ultrafast NMR technique of
SOFAST is not only very important
but is also very difficult. For instance,
reducing the recycling delay in the
SOFAST experiments means that the
duty cycle of the NMR pulse sequence
increases dramatically, from the typical
8% to over 80% (RF pulses and N15
decoupling). Imagine pushing this
increased RF load into a tiny antenna
(RF coil) that needs to be maintained
at cryogenic temperatures with a tight
precision (higher then 0.1K stability)
using a closed-cycle cryogenic refrigerator system.11 This challenge seemed
so significant that researchers at the
Institute of Structural Biology (IBS) in
Grenoble, France, have developed a
modified version of the SOFAST
experiment with no N15 decoupling,
and thus a reduced duty cycle, to
ensure that the sequence could be run
on a cryogenic probe.5
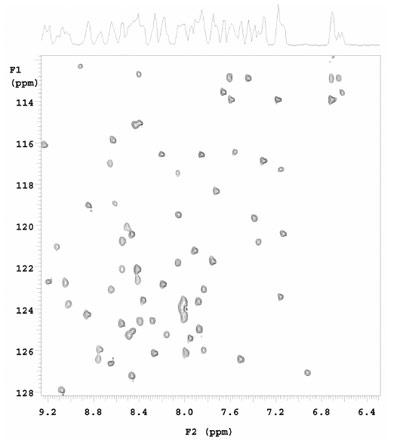
Figure 4 - SOFAST HMQC collected on N15, C13 labeled 1 mM ubiquitin (90% H2O, 10%
D2O) using a 5-mm Varian triple resonance Z-PFG cold probe. Experimental time was 8 sec. Data
courtesy of Dr. Ashish Arora (Central Drug Research Institute, Lucknow, Uttar Pradesh, India).
The rewards of being able to completely
align the cold probe technology and the
very fast SOFAST method are very significant,
and the technology was carefully
evaluated by the Varian applications and
R&D team. They have shown that on
the very robust Varian cold probes it is
possible to run the SOFAST-HMQC
experiments with N15 decoupling without
any special setup or hardware requirements.
As an example, data recently collected
on a newly installed 600-MHz Varian
cold probe at the Molecular & Structural
Biology Division, Central Drug
Research Institute (CDRI) in Lucknow,
India, are shown in Figure 4.
One can envision maintaining the
NMR “antenna” at 25K while almost
continuously bombarding it with
high-power RF pulses and decoupling,
and doing so not for a few seconds, but
for hours. This would be of particular
significance for high-throughput laboratories
that require performing series
of back-to-back 2-D experiments.
Studies at the Varian NMR laboratories
have shown that the duration of
such power-intensive experiments can
be extended considerably. Thus,
impressive power handling capabilities
of the cold probe open new
avenues for detailed studies of protein
dynamics and noticeably increase the
throughput of similar experiments.
Overall, the combined use of the
SOFAST NMR method and the
robust Varian cold probe is revolutionizing
the way researchers can study fast
protein dynamics and gain better
insight into the world of proteomics.
References
- Freeman, R.; Kupče, Ē. J. Biomol.
NMR2005, 27, 101–13.
- Malmodin, D.; Billeter, M. Prog.
Nucl. Magn. Reson. Spectrosc.2005,
46, 109–29.
- Kupče, Ē .; Nishida, T.; Freeman, M.
Prog. Nucl. Magn. Reson. Spectrosc.2003, 42, 95–122.
- Schanda, P.; Brutscher, B. J.Am.
Chem. Soc. 2005, 127, 8014.
- Schanda, P.; Kupče, Ē.; Brutscher, B.
J. Biomol. NMR 2005, 33, 199–211.
- Gal, M.; Mishkovsky, M.; Frydman, L.
J. Am. Chem. Soc., in press (2006).
- Bougault, C.; Feng, L.; Glushka, J.;
Kupče, Ē .; Prestegard, J.H. J. Biomol.
NMR2005, 28, 385–90.
- Pervushin, K.; Vögeli, B.; Eletsky, A. J.
Am. Chem. Soc. 2002, 124, 12, 898–902.
- Ernst, R.R.; Bodenhausen, G.;
Wokaun, A. Principles of Nuclear Magnetic
Resonance in One and Two
Dimensions. Oxford Science Publications:
Oxford, U.K., 1987.
- Kupče, Ē; Freeman, R. J. Magn.
Reson. 1993, 102A, 122–6.
- Losonczi, J.; Green, I. Am. Lab. 2004,
36, 26–9.
- Flynn, P.F.; Mattiello, D.L.; Hill,
H.D.W.; Wand, A.J. JACS 2000, 122A, 4823–4
Dr. Losonczi is Marketing Manager, NMR
Probes; Dr. Kupče is a Principal Applications Scientist
and Varian Fellow; and Dr. Gray and Dr.
Sandor aredor are Senior Applications Scientists, Varian
Inc., 3120 Hansen Way, Palo Alto, CA 94303,
U.S.A.; tel.: 650-424-3826; fax: 650-494-7186; e-mail: [email protected]. Dr.
Brutscher is a Research Scientist, Institut de Biologie
Structurale Jean-Pierre Ebel CNRS-CEAUJF,
Grenoble, France.