Understanding how biodiversity is distributed in nature and how it is impacted by human activities has become crucial for nature conservation. When analyzing biodiversity, it is essential to comprehend the dynamics and geographical distributions of genetic variability in natural populations. Angiosperms (or flowering plants) exhibit different modes of inheritance and patterns of genetic information among individuals. The main mechanisms generating and exploiting genetic variation are based on diverse reproductive strategies used by plants, from imposed cross-pollination to asexual reproduction via seed (apomixis) throughout different intermediate syndromes.1 Unlike sexually reproducing species, facultative apomicts can produce both sexual and asexual derived seeds through the formation of unreduced female gametophytes.2
Until a few decades ago, the use of time-consuming microsectioning techniques was the only way to learn with certainty about these reproductive processes in plants, and by then, most of the different reproductive developmental patterns currently known had already been established.3,4 Nonetheless, the arrival of new technologies has allowed us to more simply and quickly unravel the main reproductive syndromes of plants (sexual vs asexual reproduction; see Figure 1) by using special methodologies such as flow cytometry (FC).
 Figure 1 – Schematic representation of reproductive pathways in facultative apomictic species. The embryo sac structure and ploidy of cells determine the relative ratio of embryo:endosperm DNA content. In the sexual pathway (a), double fertilization leads to the formation of a 2n (2C) embryo and a 3n (3C) endosperm (see Ref. 5), whose cells can be distinguished by the relative position of the peaks in the flow cytometric histograms. In the asexual pathway (b), fertilization occurs only in the central cell of the unreduced embryo sac, and a 2n (2C) parthenogenetic embryo and 5n (5C) pseudogamous endosperm are formed, resulting in a different histogram peak configuration. em: embryo; en: endosperm (for details see Ref. 5).
Figure 1 – Schematic representation of reproductive pathways in facultative apomictic species. The embryo sac structure and ploidy of cells determine the relative ratio of embryo:endosperm DNA content. In the sexual pathway (a), double fertilization leads to the formation of a 2n (2C) embryo and a 3n (3C) endosperm (see Ref. 5), whose cells can be distinguished by the relative position of the peaks in the flow cytometric histograms. In the asexual pathway (b), fertilization occurs only in the central cell of the unreduced embryo sac, and a 2n (2C) parthenogenetic embryo and 5n (5C) pseudogamous endosperm are formed, resulting in a different histogram peak configuration. em: embryo; en: endosperm (for details see Ref. 5).Although the first patent on a prototype was issued in 1953, the first flow cytometry device based on fluorescence was developed in 1968 by Wolfgang Göhde from the University of Muenster (Germany) and was first commercialized in 1968–69 by Partec GmbH (Muenster, Germany) through PHYWE AG (Goettingen, Germany).6,7
Development of flow cytometry methods
Flow cytometry was initially developed for medical applications, but later it became useful in different fields for such applications as cell counting, sorting, and biomarker detection.7 Although the first paper on plant nuclei analysis using flow cytometry was published in 1973,8 the breakthrough in plant DNA analysis by FC came during the late 1980s, when researchers started to persistently apply this technology in plants. Basically, botanists use flow cytometry to measure DNA content in plant nuclei. For a detailed description of the principles and applications of flow cytometric cell analysis in plants, see Ref. 9.
 Figure 2 – a) Schematic overview of a typical flow cytometer technology setup (courtesy of M. Steinberg). Information from every plant nuclei is collected by different detectors, one aligned to the light beam (forward scatter), and several perpendicular to it (side scatters). b) CyFlow device (Partec GmbH) located at the Department of Systematic Botany, Albrecht-von-Haller Institute for Plant Sciences, Georg-August University of Goettingen.
Figure 2 – a) Schematic overview of a typical flow cytometer technology setup (courtesy of M. Steinberg). Information from every plant nuclei is collected by different detectors, one aligned to the light beam (forward scatter), and several perpendicular to it (side scatters). b) CyFlow device (Partec GmbH) located at the Department of Systematic Botany, Albrecht-von-Haller Institute for Plant Sciences, Georg-August University of Goettingen.Figure 2a shows an overview of how the flow cytometer lines up stained or targeted particles (sized between 0.2 and 150 μm7) by suspending them in a hydrodynamically focused stream of fluid through electronic detection equipment. In plant research, nuclei (i.e., particles) are stained with a fluorescence marker (e.g., DAPI [4’,6-diamidino-2-phenylindole] or propidium iodide). Then, each suspended nuclei is exposed to a beam of light (usually laser or UV light), after which the light path is scattered and perceived by detectors that analyze the fluorescence intensity of every individual particle, providing information about the DNA content of the nucleus. The pooled data of thousands of intact nuclei give information about the DNA content of the respective tissues (see peaks in Figure 1). Since the physical and/or chemical characteristics of up to thousands of particles per second are simultaneously analyzed, the quality and purity of solutions used for sample preparation and sample examination are crucial.
When a low-quality or inappropriate water supply system is used, the presence of contaminating particles that do not belong to the sample can exhibit fluorescence and create “noise,” interfering with the results and causing inaccurate evaluations. Hence, the use of high-quality ultrapure water is mandatory.
Ultrapure water system for production of ultrapure ASTM Class I quality water
Ultrapure water systems that produce ultrapure ASTM Class I quality water are offered by Sartorius (Goettingen, Germany). To evaluate the effects of using ultrapure water (ArUPH2O) produced by the arium® pro VF water system on the quality of the results obtained by flow cytometric analysis, the authors examined the reproductive pathways used by a facultative apomictic angiosperm to produce seeds. This was done by employing the ArUPH2O to run the samples and comparing the results to those obtained using the standard sheath fluid solution (0.04% sodium azide, 0.01% detergent) (Partec GmbH).
 Figure 3 – arium pro VF ultrapure water system (photo courtesy of Sartorius).
Figure 3 – arium pro VF ultrapure water system (photo courtesy of Sartorius).The arium pro VF system (Figure 3) produces ultrapure water from pretreated drinking water by removing all contaminant particles. Ultrapure water production requires continuous recirculation and a constant water flow rate, which is achieved using a pump system with controlled pressure. The conductivity of the water is measured at the feed water inlet and at the downstream port, or product water outlet.
The arium pro VF system used in the test described here (predecessor model with the same technical design as that of the current arium pro VF system shown in Figure 3) operates with two different cartridges. These are filled with a special active carbon adsorber and mixed-bed ion exchange resins to deliver ultrapure water with a low total organic carbon (TOC) content. Moreover, the unit has an integrated UV lamp that has an oxidizing and bactericidal effect at wavelengths of 185 nm and 254 nm, respectively.
 Figure 4 – Schematic flow diagram for the arium pro VF ultrapure water system (for enhanced clarity, the valves and their controllers are omitted).
Figure 4 – Schematic flow diagram for the arium pro VF ultrapure water system (for enhanced clarity, the valves and their controllers are omitted).The ultrapure water system has a built-in ultrafilter module used as a crossflow filter. The ultrafilter membrane utilized in this filter retains colloids, microorganisms, endotoxins, RNA, and DNA. A 0.2-μm final filter installed at the water outlet serves to remove particulates and bacteria during dispensing of the ultrapure water stream produced. The process employed by the unit to produce ultrapure water is shown in Figure 4.
Materials and methods
Seed samples from hexaploid Ranunculus carpaticola plants were collected under open pollination conditions. Individual seeds were chopped in a plastic petri dish with 300 μL of extraction buffer (CyStain UV Precise P, Partec GmbH) and incubated for 10 min at room temperature, after which nuclei were filtered (30 μm mash, CellTric®, Partec GmbH) into a 5-mL plastic tube. Then, 1.2 mL of staining buffer (CyStain UV Precise P) was added, and after 60 sec, samples were analyzed in a flow cytometer (CyFlow Space, Partec GmbH; see Figure 2b) in the blue fluorescence channel.
 Figure 5 – Flow histograms of hexaploids
Figure 5 – Flow histograms of hexaploids R. carpaticola
seeds. The x
-axis represents the relative fluorescence of every measured particle, while numbers of particles are represented on the y
-axis. a) Histogram of an apomictically derived seed exhibiting an Em:En peak ratio value of 3.15. b) Histogram of a sexually derived seed with an Em:En peak ratio value of 1.48.Setting parameters were adjusted using known diploid (2×) or tetraploid (4×) Ranunculus sp. genotypes. The relative fluorescence of at least 2000 individual nuclei per sample was quantified and represented in a histogram. The relative positions of the peaks were used to determine ploidy levels of embryo and endosperm tissues. Reproductive pathways followed during seed formation were reconstructed according to Ref. 10.
To flow the samples, two different fluids were used: A) sheath flow for flow systems (Partec GmbH), and B) ultrapure water from arium pro VF. Statistical analyses for trial comparisons were conducted with an Excel spreadsheet (Microsoft Corp., Redmond, WA).
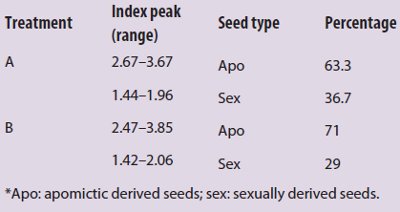
Table 1 – Reproductive classification of R. carpaticola seeds according to the embryo:endosperm peaks ratio*
Results
The reproductive pathway was reconstructed for a total of 61 individual seeds. Thirty seeds were flowed using sheath fluid, and 31 seeds were flowed using ArUPH2O (conductivity: 0.055 μS/cm or 18.2 MΩ × cm resistivity compensated to 25 °C) following the standard recommended procedure. An average of 2329 nuclei were measured per sample, representing 73% of the total particles counted, while the other 27% was represented by nuclei in the G2 phase of cell cycle and background signals. Embryo and endosperm mean peak positions were calculated for all analyzed seeds based on an averaged total number of nuclei gathered for every peak.
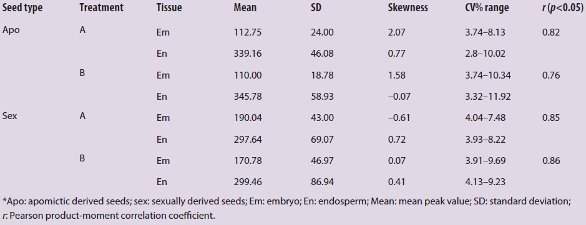
Table 2 – Statistical analyses of flow cytometry histograms from 61 R. carpaticola seeds*
Relative positions of nuclei on an x-axis histogram plot are based on its relative ploidy according to the tissue of origin (see Figure 1). Hence, a tetraploid nucleus (4n) will have approximately twice as much DNA as a diploid one (2n), emitting a twofold intense fluorescence signal and therefore being placed at twice the relative distance on the x-axis (Figure 5). The reproductive origin of each seed and proportion of sexuality and apomixis were established after index peak determination (Table 1).
Different statistical parameters were taken for each trial and reproductive pathway to compare treatments (Table 2). Absolute values of the coefficient of variation (CV%) were obtained for every sample, and ranges per trial and reproductive pathway, as well as the skewness values of each embryo and endosperm-tissue peak for all apomictic and sexual samples, are presented in Table 2. The Pearson product-moment correlation was calculated for mean values of embryo and endosperm peaks measured per sample and fluid treatment (Table 2).
Discussion
The hexaploid cytotypes of Ranunculus carpaticola are facultative apomictic plants that use a variety of reproductive strategies to generate new individuals in natural populations.11 Flow cytometric seed screening is a useful, easily applied, and timeless technique for the evaluation of reproductive strategies in seed plants. The two independent treatments employed here (sheath fluid vs ArUPH2O) do not show significant differences in peak quality, peak position, or index peak ranges.
The coefficient of variation is a normalized measure used to quantify how dispersed or clustered a set of observed events is compared to a standard probability distribution or frequency distribution (an assumed statistical model). The absolute values of the CV or relative standard deviation (RSD or %RSD) correspond to CV values expressed as a percentage. In this case, though maximum CV% values were slightly higher when using ArUPH2O, the difference to the sheath fluid treatment was always smaller than the minimum CV% values, pointing to a similar level of dispersion of counts between treatments.
In addition, even though the bell-shaped peaks tend to have longer right tails (positive skewness), the values are close to zero, indicating the peak had a nearly symmetric shape around the mean. Moreover, the high correlation observed between values of embryo and endosperm peaks indicates that the use of ArUPH2O does not affect the mean relative intensity of measured fluorescence when evaluating nuclei populations, thus preventing distorted correlations and maintaining a dispersion of values within expected ratios for embryo-to-endosperm peaks.
However, proportions of sexual to asexual seeds were similar in both treatments, as expected when randomly selected samples are used. These results indicate that the determination of reproductive origins is not biased by unproportional shifting of peak positions between treatments.
Overall, comparatively few discrepancies in the obtained results prove the arium system is a time-effective and economical alternative for the analysis of relative ploidies in plant materials. The high quality of arium particle-free water proved to be effective in providing reproducible results in flow cytometric seed screenings. However, since additives like antibiotics and detergents have positive short- and long-term effects on the reliable operation of the flow cytometry system by preventing biofilm formation and improving wetting of the inner surfaces of containers, tubing, valves, and flow cells, the supplier recommends the addition of such substances to a self-made sheath fluid.
To summarize, ArUPH2O can be readily used for flow cytometry of plant cells. Because flow cytometric technology is gaining importance in other applications, such as detection of tumor cells, quantitative estimation and morphological differentiation of cells, cell cycle analysis, DNA–RNA content estimations, and apoptosis measurements, the suitability of ArUPH2O is opening up new opportunities for arium ultrapure water in a variety of applications using the emergent technology of flow cytometry.
References
- Johri, B.M. Embryology of Angiosperms. Springer-Verlag: Berlin, Germany, 1984.
- Nogler, G.A. Gametophytic Apomixis. In: Embryology of Angiosperms; Johri, B.M., Ed. Springer-Verlag: Berlin, Germany; pp 475– 518.
- Battaglia, E. The male and female gametophytes of angiosperms—an interpretation. Phytomorphology 1951, 1, 87–116.
- Asker, S.E.; Jerling, L. Apomixis in Plants. CRC Press: Boca Raton, FL, 1992.
- Hojsgaard, D.H.; Martínez, E.J. et al. Competition between meiotic and apomictic pathways during ovule and seed development results in clonality. New Phytologist 2013, 197, 336–47 (and supporting information in the online version of this article).
- Dittrich, W.; Göhde, W. Patent DE 1815352, Flow-through Chamber for Photometers to Measure and Count Particles in a Dispersion Medium; 1968.
- http://en.wikipedia.org/wiki/Flow_cytometry.
- Heller, F.O. DNS-Bestimmung an Keimwurzeln von Vicia faba L. mit Hilfe der Impulscytophotometrie. (DNA estimation on radicles of Vicia faba L. using pulse cytophotometry; translation of the original German title by Dr. Herbig.) Berichte der Deutschen Botanischen Gesellschaft 1973, 86, 437–41.
- Doležel, J. Flow cytometric analysis of nuclear DNA content in higher plants. Phytochem. Anal. 1991, 2, 143–54.
- Matzk, F.; Meister, A. et al. An efficient screen for the reproductive pathways using mature seeds of monocots and dicots. Plant J. 2000, 21, 97–108.
- Paun, O.; Hörandl, E. Evolution of hypervariable microsatellites in apomictic polyploid lineages of Ranunculus carpaticola: directional bias at dinucleotide loci. Genetics 2006, 174, 387–98.
Dr. Diego H. Hojsgaard is with the Albrecht-von-Haller Institute for Plant Sciences, Department of Systematic Botany, Georg-August University, Untere Karspuele 2, 37073 Goettingen, Germany; tel.: +49 (0) 551/39 5731; fax: +49 (0) 551/39 22329; e-mail: [email protected] . Dr. Elmar Herbig is with Sartorius AG, Goettingen, Germany. The authors acknowledge Matthias Steinberg (Partec GmbH, Muenster, Germany) for his helpful discussions and suggestions, and for providing the flow cytometry setup image (Figure 2a). The authors also thank Prof. Dr. Elvira Hörandl and Dr. Simone Klatt (Department of Systematic Botany, Georg-August University, Goettingen, Germany) for their critical reading of the manuscript.