It is essential to eliminate poor drug candidates as early as possible in the drug development process. Unfavorable pharmacokinetic (PK) properties have frequently been the reason for failure of new chemical entities (NCEs) in the clinic. This realization led to the application of high-throughput principles to the in vivo lead optimization process, wherein the NCE is first administered to rodents to determine the PK profile in a rapid and limited study. This rapid primary screen is known as Fast PK (Taylor Technology, a subsidiary of PharmaNet Development Group, Inc., Princeton, NJ), and several critical decisions are based on this initial study.
While triple-quadrupole systems have been dominant in pharmaceutical bioanalysis, recent fundamental advances in MS technology have resulted in a mass analyzer called the Orbitrap™ (OT) (Thermo Fisher Scientific, San Jose, CA). Much like a triple-quadrupole, OT technology can enhance the lead optimization process by providing PK data for the NCE, but with the added capability of providing simultaneous, semiquantitative information on metabolites present in the incurred sample. This allows informed decisions to be made further upstream in the discovery process, thereby reducing the cost of new drug development.
Impact of MS on Fast PK
The terms “cassette dosing PK” (NCE cocktail dosed per animal), “cassette accelerated rapid rat PK” (one NCE per animal, 24 hr), “Fast PK” (one NCE per animal, 8 hr), and “snapshot PK” (one NCE, mouse model, 3 hr) became the norm for improving in vivo cycle time for lead optimization studies in large pharmaceutical companies. These approaches are important because they reduce the number of animals involved; the number of samples to be analyzed; and, in many cases, both. In addition, for a large majority of cases, rodent Fast PK studies correctly predict human PK, thus increasing the success rate of the NCE as it moves down the pipeline.
A basic Fast PK rodent assay includes formulation, dosing, and bioanalysis (plasma) of six time points in six rats (3 intravenous [IV] and 3 oral [PO]) in 6 hr. The full PK rodent assay would extend the number of time points to 8 and out to 24 hr postdose. There are several advantages to this approach: 1) It provides information to the synthetic chemist so that the chemical structure of the NCE may be fine-tuned, and 2) it enables better design of in vivo efficacy studies benefiting PK observations. Essentially, the Fast PK study validates oral bioavailability (solubility) and intestinal absorption (permeability) characteristics, the two key parameters indicative of future success.
Fast PK quantitative studies are traditionally performed using a technique known as selected reaction monitoring (SRM) on a triple-quadrupole mass spectrometer. In the SRM mode, the ions specific to the NCE are focused onto the detection system, while all other nonspecific ions are filtered away. As a result, only the NCE is quantified and its PK estimates determined, but any metabolite information contained in the sample is lost. With the understanding of this limitation of the SRM technique is the acknowledgment that potentially efficacious NCEs may be prematurely eliminated due to poor PK characteristics, when in fact more complete feedback from the same Fast PK study could provide the support for the decision to make a simple tweak of the chemical structure to improve bioavailability. OT is able to provide metabolite information during the Fast PK analysis.
Ease of use is the fundamental driver for utilization of high-resolution mass spectrometry (HRMS) for quantitative bioanalysis. It can be performed without any prior knowledge of the compound, thus avoiding the need for compound-specific tuning (setting up the SRM transition). This has significant ramifications in Fast PK applications, especially since biotransformation information can now be provided. The Exactive Orbitrap can quantify both the NCE and its major biotransformation products simultaneously, using high resolution and accurate mass capability to filter away chemical noise and capturing more information per scan. Triple-quadrupole MS instruments usually have a resolution value in the few hundreds, whereas the Exactive Orbitrap operates at a nominal value of 10,000; this can be increased to 100,000, if needed.
When using HRMS, compound-specific MS-MS tuning is no longer required for specificity; very high resolution (R>10,000) in full MS mode is sufficient. Since the fragmentation process intrinsically reduces sensitivity, only S/N is gained; HRMS full MS mode applications theoretically do not have this limitation. Whereas HRMS simply comes down to being dependent on the ionization efficiency, the SRM scan can be viewed as the sum of the ionization efficiency and fragmentation efficiency. Tuning several SRM transitions daily in a discovery setting is time consuming, and automatic tuning algorithms have been developed. The OT’s accurate mass functionality enables elemental composition information, allowing identification of biotransformation products (metabolites), while it is performing Fast PK bioanalysis.
Bioanalysis and semiquantitative analysis of metabolites using HRMS
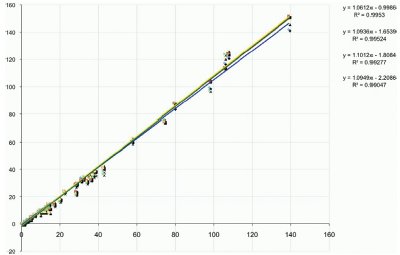
Figure 1 - Correlation between triple-quadrupole versus Exactive bioanalysis at different resolution settings (R = 10K, 25K [2×], and 50K).
Figure 1 shows a comparison between bioanalysis results from a triple-quadruple (TSQ Quantum Ultra, Thermo Fisher Scientific, Austin, TX) and the Exactive Orbitrap. A perfect correlation would indicate a slope of 1.000, and the quantitative performance is almost indistinguishable between the two instrument platforms (see Figure 2) for discovery applications where the lower limit of quantitation (LLOQ) is usually between 1 and 5 ng/mL.
TSQ Exactive
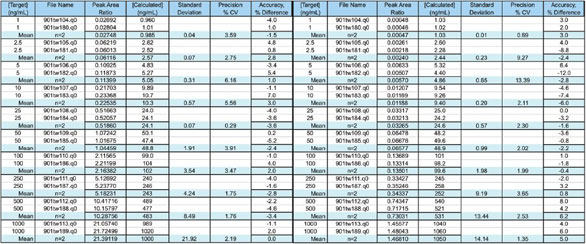
Figure 2 - Bioanalysis data in the range 1–1000 ng/mL. Data from TSQ (left) and Exactive (right) meet the LOQ criteria of <20% for accuracy and precision, indicating that either platform can be used for bioanalysis.
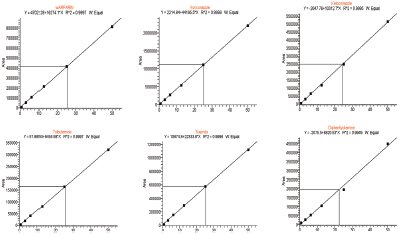
Figure 3 - Examples of 6 out of 17 compounds quantified in full-scan MS mode on the OT using 25K (2-ppm window).
To further prove the quantitative performance using HRMS, 17 commercially available compounds were quantified in rat plasma in the range 1–50 ng/mL (see Figure 3) without tuning of any parameters. This is important because one key application of the OT platform in discovery applications is the semiquantitative analysis of metabolites during the first in vivo PK study. To test this hypothesis, chlorprazomine was incubated with microsomes for 60 min, and the metabolites were identified and quantified without any a priori knowledge on the Exactive OT platform.
The key to identifying metabolites “on-the-fly” using HRMS is utilizing the advantage provided by mass defect, which is the difference between the exact mass and the integer mass (equivalent to the number of protons and neutrons) as a result of mass deficiency. This mass deficiency is due to the fact that 12C is the only isotope with an integer exact mass of 12.00000 (ad infinitum) and used arbitrarily to set the atomic mass scale for all other elements. Therefore, 16O has an isotopic mass of 15.9949 u, and is deficient by 5.085 mmu. Interestingly, 1H precedes 12C and therefore weighs 1.0078 u, gaining almost 7.825 mmu per 1H. Since 1H is ubiquitous, a typical small molecule containing 40 hydrogen atoms can lead to a positive mass defect of about +0.3 Da.
A positive mass defect is also referred to as mass sufficiency as opposed to mass deficiency, typically exhibited by elements with mass numbers higher than 12C on the periodic table (14N, 16O, 32S, 35Cl). Instruments such as the OT can also determine the mass difference between the 1H (1.007824 Da) and a proton +H (1.007276 Da), which is deficient of 1 electron or 0.000548 Da. This becomes a factor for the analysis of peptides that are multiply charged (z = n, where n>1), and the difference in mass can be used to advantage.
Table 1 Abbreviated list of common phase I and phase II metabolites with their mass defects, added to the parent compound m/z generated in MS Excel*
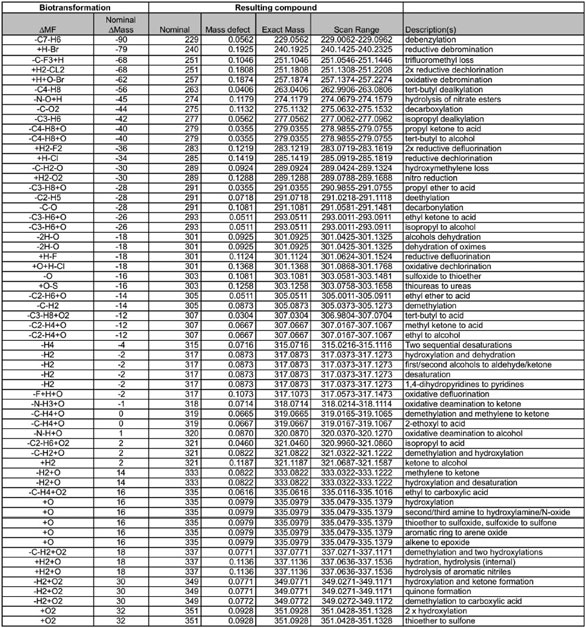
*Microsoft (Redmond, WA). Once the elemental formula for the parent NCE is entered, an automated, accurate mass list (with mass defects) is generated. The acquired data file can then be queried (postacquisition) using this Excel-generated list.
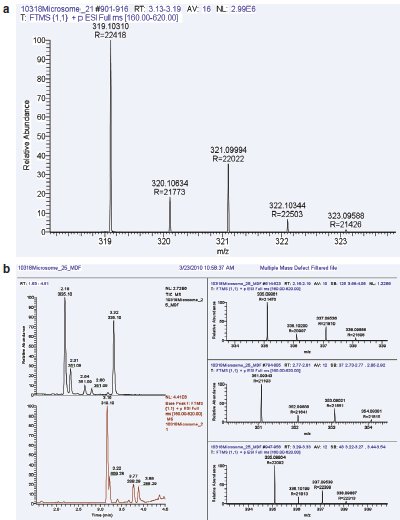
Figure 4 - a) HRMS (R = 20,000) mass spectra of chlorprazomine. b) In vitro generation of metabolites (microsomal incubation, 60 min) of chlorprazomine (retention time, 3.19 min). Top panel shows the mass defect filtered chromatogram, indicating the formation of three abundant metabolites of chlorprazomine (ret. time: 2.18, 2.31, 2.64, 2.80, 3.32 min) with full-scan mass spectra (right panel) showing 2 ppm or less mass accuracy. The Cl isotope pattern can also be used to potentially identify the metabolites, as can be seen in the mass spectra.
Table 1 shows that, since the common phase I and phase II metabolites have their own mass defects, they become additive to the precursor ion upon biotransformation. Thus, their masses can simply be added or subtracted from the exact measured mass of the precursor ion, and metabolites can be quickly identified and then quantified (see Figure 4). The presence of a characteristic isotopic pattern, in this case chlorine, helps in the potential identification of metabolites substantiating mass defect-based filtering (see Figure 5). The quantitative analysis is performed against the precursor and, since the response factors for the metabolites are not available at this early stage, the quantitative analysis is, at best, semiquantitative or relative to the parent. This is an exciting new application of HRMS for drug discovery applications to further improve the viability of the NCE under study, where metabolism information can be provided to the synthetic chemist as well as the pharmacologist.
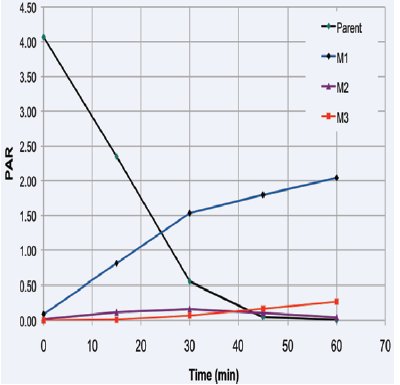
Figure 5 - Semiquantitative analysis of chlorprazomine (m/z 319.1030) parent and its metabolites.
For example, the rapid formation of an active hydroxy metabolite (+15.9949) can provide valuable information in terms of potential bioavailability, despite poor oral PK profile, especially during the first in vivo PK study where the NCE is dosed both orally and via IV. The formation of a hydroxy metabolite for chlorprazomine at m/z 355.0979 (M1 or m/z 319.1030 + 15.9949 = 335.0979) is quantified in Figure 4; MS-MS data will yield the exact location on the molecule (soft spot), but this is outside the scope of this study. The common β-blocker, alprenolol, for example, based on its plasma concentration, is more active given a single oral dose than when given via IV, despite exhibiting poor oral bioavailability in terms of PK. Alprenolol forms an active 4-hydroxyalprenolol metabolite responsible for therapeutic action. Using HRMS, this information can now be available in the very first in vivo study, compared to the traditional SRM technique, which focused only on the precursor ion (NCE only) and is blind to all other information (metabolites) in the incurred sample.
Conclusion
LC-MS provides the absorption, distribution, metabolism, and excretion (ADME) research scientist with a powerful analytical tool, and has played a major role in drug discovery applications. Over the past 20 years, bioanalysis has been performed using SRM on a triple-quadrupole with a singular focus on small molecules. However, it is likely that with the emergence of biopharmaceuticals and the routine ease of use offered by time-of-flight (TOF) and OT MS, these techniques will attain the sensitivity and specificity required of nonregulated bioanalysis, perhaps signaling a paradigm shift.
Dr. Thakur is Associate Director, Specialized Mass Spectrometry, and Mr. Koleto is Senior Scientist, Taylor Technology, a subsidiary of PharmaNet Development Group, Inc., 504 Carnegie Center, Princeton, NJ 08540, U.S.A.; tel.: 609-951- 6800; fax: 609-514-0390; e-mail: [email protected].