In vivo protein imaging technology
has been greatly advanced over the last
decade by the development of green fluorescent
protein (GFP) and its derivatives
as investigational tools. Typical applications
of GFP involve tagging a protein of
interest so that the protein expression and
localization can be imaged as the fusion
protein is being expressed in live cells.
In contrast to protein imaging, RNA imaging
in live cells lags far behind. The currently
available in vivo RNA imaging methods
are 1) fluorescence in situ hybridization,1,2
involving fluorescently labeled oligonucleotides
as probes that bind to complementary
RNA sequences in fixed cells; 2) a molecular
beacon, an alternative oligonucleotide-based
probe labeled with a fluorophore and
a quencher at each end;3,4 3) a protein-based
RNA biosensor system that requires that GFP
be fused to the RNA-binding protein MS2
and that the RNA of interest be linked to
many copies of MS2-binding RNA sequence;5
and 4) the quenched autoligating fluorescence
resonance energy transfer (FRET) probes.6 All of these
methods involve the use of biopolymers as fluorescent
probes, which may be a major concern since biopolymers
are prone to rapid degradation.
New RNA imaging technology would significantly
enhance our understanding of rarely investigated
transcription kinetics and RNA trafficking in live
cells. The focus of this article is the development of
a new paradigm for RNA imaging technology. The
long-term goal of the research program described is to
tag an RNA of interest with a fluorescence-inducing
RNA aptamer (reporter RNA) at the DNA level and
investigate its transcription and RNA trafficking using
RNA chemosensors (small molecules that report specific
RNA) in live cells. Analogous to GFP, this fluorescence-inducing aptamer will be used universally as
a tagging (investigational) tool to study various genes.
RNA imaging with small
molecules
A desirable RNA sensor should be cell permeable,
stable, and fluorescent only when bound to specific
RNA. Some of the benefits of such a chemosensor
are as follows:
-
Organic synthesis allows for tuning chemical and
photophysical properties of chemosensors
- Chemosensors enable better quantification of
RNA than biosensors due to the linear correlation
between the emission signal and quantity of
RNA–sensor complex
- Cell-permeable RNA chemosensors allow for
both noninvasive introduction of the molecule
to live cells and animals and microinjections to a
specific area inside cells
- Chemosensors are less likely to be degraded in
live cells than biosensors, permitting longer time-scales
for live cell imaging; of particular interest
is the potential use of an RNA chemosensor to
monitor the dynamics of RNA concentrations
during apoptosis, which presumably cannot be
studied using currently available technology
- The stability and cell permeability of chemosensors
provide a more steady concentration in biological
samples
- Chemosensor-bound RNA is expected to behave
more similarly to natural RNA than biomacro-molecule-bound RNA.
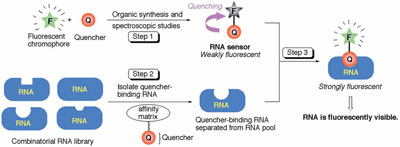
Figure 1 - Overall strategy for developing reporter RNA systems.
Figure 1 illustrates a new paradigm for small-molecule-based
RNA sensors developed in the University of Pittsburgh
Department of Chemistry laboratory (Pittsburgh,
PA).7 In step 1, using synthetic organic chemistry, a
quencher (Q) is covalently linked to a fluorophore (F) to
provide a fluorogenic compound (F-Q). By virtue of the
photoinduced electron transfer (PET; for a description of
PET, please see below) process from Q to F, the fluorescence
of F-Q is quenched. In step 2, RNA aptamers for Q
(but not for F) are raised and isolated by means of in vitro
RNA selection. In step 3, fluorescence spectroscopic
analysis of the aptamer–F-Q mixture is performed; these
aptamers suppress the quenching function of Q, thereby
restoring the fluorescence signal from F.
The physical chemistry
underlying the
paradigm: PET
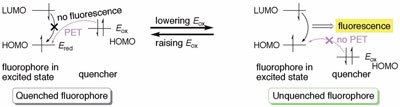
Figure 2 - Ability of PET to quench fluorescence.
A desirable probe fluoresces when bound to a specific
biomolecule, while generating no signal when
unbound. This fluorescence on-off switch can be
implemented by the PET mechanism
(Figure 2).8 When a quencher is proximate
to a fluorophore and its highest
occupied molecular orbital (HOMO)
energy level is sufficiently high (Figure
2, left), PET occurs from the quencher to
the excited fluorophore without fluorescence
emission. According to the Rehm-Weller equation (Eq. [1]),9 lowering Eox
increases the fluorophore’s fluorescence
signal (Figure 2, right). This theory is
well-established10 and has been used to
rationally design chemosensors for biologically
important ions.11–16
ΔGPET = Eox – Ered – ΔE00 – wp (1)
where Eox = oxidation potential of a
quencher, Ered = reduction potential of
a fluorophore, ΔE00 = the singlet excited
energy, and wp = the work term for the
charge separation state.
Fluorescence quenching is controlled not
only by potential energy but also by the
concentration of quencher as indicated by the Stern-Volmer equation (Eq. [2]). This equation shows that
as [Q] increases, fluorescence quantum yield Φ drops
dramatically. This was taken into account and it was
decided to append two quenchers to a fluorophore.
Φ0/Φ = (1 + KD[Q]) (1 + Ks[Q]) (2)
where Φ0 and Φ are the quantum yields in the
absence and presence of quencher, respectively; KD
and KS are the Stern-Volmer constants of dynamic
and static quenching, respectively; and [Q] is the
concentration of quencher.
Results and discussion
Synthesis of 2′,7′-dichlorofluorescein
derivatives
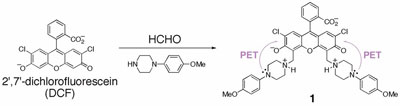
Figure 3 - Synthesis of compound 1 from DCF.
As part of step 1 (Figure 1), commercially available
2′,7′-dichlorofluorescein (DCF) was converted to compound
1 and others (not shown) in one step (Figure 3),
and their quantum yields were determined.17 From these
studies, compound 1 emerged as a potential sensor due to
its lowest background emission (Φ1 = 0.025) presumably
by the PET process from aniline nitrogen atoms.
In vitro RNA selection and
fluorescence analysis
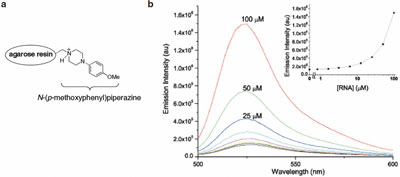
Figure 4 - a) Structure of affinity column. b) Concentration-dependent fluorescence induction
of compound 1 (1 μM) with its aptamer (graphs taken from the literature). (Reproduced with permission
from Ref. 7.)
The approach depicted in Figure 1 is unique because
it requires RNA that binds to a quencher and not to a
fluorophore. To identify such aptamers, the quencher of
1, N-(p-methoxy-phenyl)piperazine was linked to agarose
resin (Figure 4a).7 The resulting matrix was used as bait
in a series of in vitro RNA selections using an N70 RNA
library (70 nucleotides were randomized), and three RNA aptamers were characterized.7 It was gratifying
to see that one of the three RNA aptamers enhanced
the fluorescence of compound 1 in a concentration-dependent
manner (Figure 4b). The addition of N-(p-methoxyphenyl)
piperazine was found to antagonize
this fluorescence induction effect, suggesting that the
aptamer induces the fluorescence of compound 1 by the
mechanism envisioned (Figure 1, step 3). The mixture
of the N70 RNA library did not enhance fluorescence,
excluding nonspecific fluorescence enhancement by
RNA. Despite the high concentrations of RNA used in
the titration experiments (20 μM to induce threefold fluorescence
enhancement), this result provides the proof
of concept for the approach illustrated in Figure 1.7