Lactase (β-D-galactohydrolase EC 3.2.1.23) is widely distributed in microorganisms, plants, and animals.1 Lactase is a hydrolytic enzyme that hydrolyzes the terminal nonreducing β-D-galactoside residues from β-galactosides to glucose and galactose, which are simple monosaccharides. It has been exploited extensively in the pharmaceutical and food industries.2,3 It is widely used in the pharmaceutical industry to prevent problems of lactose intolerance by individuals who are lactase deficient.4
The use of lactase for the hydrolysis of lactose in milk and whey is one of the most important applications in the food industry.5 There is also considerable interest in the use of lactase in processing foods containing galactooligosaccharides to inhibit sucrose crystallization from beet sugar syrups.6 The hydrolysis of galactooligosaccharides may alleviate gastric distress caused by fermentation of carbohydrates in the large intestine;7 therefore the improvement of the nutritional quality of legume-based foods such as soybean milk and cowpea meal can be achieved using lactase.8–10
It is well known that the properties of lactase differ markedly depending on the source. Hence, there is a need to explore new isolates for lactase production. Furthermore, most purification protocols employ two or more stages of column fractionation involving techniques such as gel filtration, ion exchange, and affinity chromatography.10–12 Multistep purification is time consuming and costly, and generally results in low yields. Single-step purification methods using affinity ligands have also been reported.13,14 However, affinity purification techniques are usually expensive, since they require specific affinity matrices to be prepared.
Hydrophobic interaction chromatography (HIC) would become more economical for lactase purification. The technique has not been fully explored, especially for the purification of lactases. The present investigation employs HIC for the purification of lactase from a new isolate of L. acidophilus.15 The Diaion HP-20™ column (Mitsubishi Kasei, Tokyo, Japan) used in this study has not been reported for lactase purification. A 110-fold purity increase and 1069 U/mg specific activity were achieved with a yield of 94.7%. The purified enzyme was further characterized.
Materials and methods
Microorganism and materials
L. acidophilus15 isolated in the authors’ laboratory was used for lactase production. Molecular weight markers (PMW-H) were from Bangalore Genei (Bangalore, India). Spectrophotometric measurements were carried out on a SPECTRONIC® GENESYS™ (Thermo Electron Corp., West Palm Beach, FL). Chromatographic experiments were done on a liquid chromatography system (Protein Purification System, Bio-Rad Laboratories, Fullerton, CA).
Determination of enzyme activity, protein, and kinetic constants
An assay of lactase activity was carried out using o-nitrophenyl galactopyranoside (ONPG). One unit of enzyme activity (U) is defined as μmol of o-nitrophenol formed per mL per min. Km and Vmax were calculated using the Lineweaver-Burk plot. The Bradford method was used for protein estimation employing bovine serum albumin (BSA) as the standard. The purity of lactase was checked by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) with silver staining.
Purification of lactase
The cells were separated by centrifugation and lysed using homogenization at 2000 psig. The supernatant after centrifugation was used for purification. This supernatant was concentrated approx. five times by ultrafiltration utilizing polysulfone membrane (nominal molecular weight cutoff [NMWC]—100 kDa). The solution was passed through a 0.45-μm filter to make it particle free. This particle-free, clear solution was loaded onto the HIC column. The NaCl stability of the enzyme was monitored for three days. NaCl was used in HIC instead of ammonium sulfate since it is inexpensive and because ammonium sulfate contains sulfite ions, which may cause impurity in the final product. One milliliter of this solution containing 2.0 M NaCl was applied onto the Diaion HP-20 column (9 mm × 98 mm) preequilibrated with 2.0 M NaCl in 0.1 M phosphate buffer, pH 7. The flow rate was maintained at 0.6 mL/min, and fractions of 3.0 mL were collected. After passing three bed volumes of the buffer, the enzyme was eluted with a linear gradient of NaCl from 2.0 M to 0 M in 0.1 M phosphate buffer, pH 7.0. The purified enzyme was subjected to electrophoresis to determine homogeneity.
Results and discussion
Lactose intolerance is a major concern in the pharmaceutical and food industries. Most work reports on the multistep purification protocol for lactase. This paper discusses the single-step purification of lactase using HIC Diaion HP-20 resin. Lactase was produced using a new isolate of L. acidophilus that was isolated from fermented Eluesine coracana.15 Since HIC is simple and efficient , it can be readily adapted for the large-scale purification of lactases. The possibility of regeneration, reuse, and mechanical stability of the matrix makes HIC a valuable industrial approach for purifying lactase in one step. The matrix used in this study has not previously been reported for lactase purification.
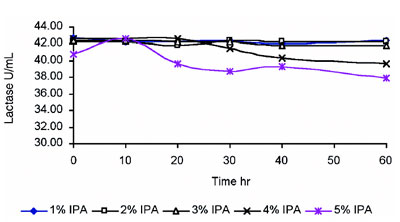
Figure 1 - Lactase stability in IPA (hydrophobic media).
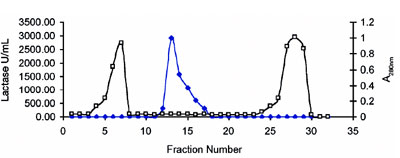
Figure 2 - Elution profile of lactase on Diaion HP-20 HIC resin using gradient NaCl elution.
Lactase was subjected to stability testing under various concentrations of isopropyl alcohol (IPA). This is required before employing HIC. The enzyme was found to be stable up to 3% IPA, which is very good stability for biologicals (Figure 1). Optimization of adsorption of the enzyme onto the Diaion HP-20 column for HIC was studied at different concentrations of NaCl, ranging from 0.0 M to 2.5 M in phosphate buffer, pH 7.0. The range of NaCl was further optimized, and 2.0 M concentration gave the best results. Similar studies were carried out to optimize the pH for purification; pH 7.0 was found to be the most appropriate. Purification of the enzyme was thus carried out in 2.0 M NaCl, pH 7.0 buffer. An elution profile of the proteins (A280 nm) showed that the majority of the contaminant proteins were eluted in the initial and end stages (Figure 2). The enzyme was eluted using a decreasing salt gradient. Lactase was eluted in column fractions 11–18 at an NaCl concentration of around 1.5 M, revealing good binding with the matrix (Figure 3). The A280 of the fractions exhibited insignificant protein in the fractions with lactase activity (Figure 2), indicating a high degree of purification. The enzyme was purified approx. 110-fold with a 94.7% yield and specific activity of 1069 U/mg protein, as shown in Table 1. Fold purity and recovery by HIC were thus superior to those obtained in most of the literature.16–18 A flow rate up to 0.6 mL/min could be employed, since Diaion HP-20 beads offer good flow properties to the medium. HIC was therefore found to save time and to be useful for processing large volumes.
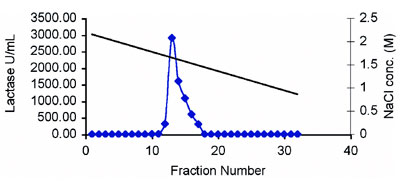
Figure 3 - Elution profile of lactase using NaCl gradient.
The purification fold and yield achieved in this study are not only comparable, but better than the results of other enzymes reported to be purified using an HIC matrix. One hundred percent adsorption of lactase on Diaion HP-20 at a much lower concentration of 2.0 M was observed in the present work. In HIC, ammonium sulfate is generally used as a salt. It is noteworthy that in the present investigation, instead of ammonium sulfate, sodium chloride was used because ammonium sulfate leaves behind the impurity of sulfite ions, which is not acceptable in pharmaceuticals. Use of NaCl imparts no impurity to the final product and also makes the process less expensive.
Lactase was purified to homogeneity. Using PAGE, the purified preparation gave a single band (Figure 4). The molecular weight of the purified L. acidophilus lactase was 400 kDa. In order to characterize the lactase, Vmax and Km were estimated using ONPG as a substrate from a Lineweaver-Burk plot. These values were 4.94 min–1 and 0.11 mM, respectively. The pH and temperature optima were observed to be 6.5 and 50 °C, respectively (results not shown).
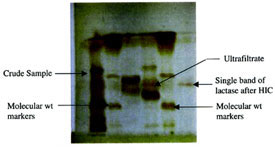
Figure 4 - Gel electrophoresis (SDS-PAGE) of lactase.
Conclusion
Lactase was produced from a new isolate of L. acidophilus and was found to have a Vmax and Km of 4.94 min–1 and 0.11 mM, respectively. The pH and temperature optima for lactase activity were 6.5 and 50 °C, respectively. The molecular weight of the purified lactase was 400 kDa. A single-step purification method for lactase from a new isolate of L. acidophilus was developed using Diaion HP-20 HIC resin. A 110-fold purity increase in lactase having a specific activity of 1069 U/mg was achieved with a yield of 94.7%. The new strain will become an important addition to science and industry.
References
- Nevalainen, K.M.H. Induction, isolation, and characterization of Aspergillus niger mutant strains producing elevated levels of β-galactosidase. Appl. Environ. Microbiol. 1981, 41, 593–6.
- Belem, M.A.F.; Lee, B.H. Production of bioingredients from Kluyveromyces marxianus grown on whey: an alternative. Crit. Rev. Food Sci. Nutr.1998, 38, 598–656.
- Bhowmick, T.; Johnson, M.C.; Bibek, R. Factors influencing synthesis and activity of β-galactosidase in Lactobacillus
acidophilus. J. Ind. Microbiol. 1987, 2, 1–7. - Gekas, V.; Lopez-Leiva, M. Hydrolysis of lactose: a literature review. Process Biochem.1985, 20, 2–12.
- Kumar, V.; Ramakrishnan, S.; Teeri, T.T.; Knowles, J.K.C.; Hartley, B.S. Saccharomyces cerevisiae cells secreting
an Aspergillus niger β-galactosidase grown on whey permeate. Bio/Technology 1992, 10, 82–5. - Kobayashi, H.; Suzuki, H. Studies on the decomposition of raffinose by α-galactosidase of mold. J. Ferment. Technol. 1972, 50, 625–32.
- Cristofaro, E.; Mottu, F.; Wuhrmann, J.J. Involvement of the raffinose family of oligosaccharides in flatulence. In Sipple, H.L., McNutt, K., eds. Sugars in Nutrition. Academic Press: New York, NY, 1974; 31,336.
- Somiari, R.I.; Balogh, E. Properties of an extracellular glycosidase of Aspergillus niger suitable for removal of oligosaccharides from cowpea meal. Enzyme. Microb. Technol.1995, 17, 311–16.
- Cruz, R.; Park, Y.K. Production of fungal α-galactosidase and its application to hydrolysis of galactooligosaccharides
in soybean milk. J. Food Sci.1982, 47, 1973–5. - Luis, A.; Harry, W.J. In flow activation of diol–silica with cyanogen bromide and triethylamine for preparing high-performance affinity chromatographic columns. J. Chromatogr. A 2003, 984, 9–17.
- Nagy, Z.; Kiss, T.; Szentirmai, A.; Biro, S. β-galactosidase of Penicillium chrysogenum: production, purification, and characterization of the enzyme. Prot. Express. Purificat. 2001, 21, 24–9.
- Amedeo, V.; Sergio, P. Separation and characterization of three β-galactosidases from Bacillus circulans. Biochim. Biophys. Acta. 1998, 1380, 223–31.
- Clemit, R.H.; Chase, H.A. Immobilized metal affinity chromatography of β-galactosidase from unclarified Escherichia coli homogenates using expanded bed adsorption. J. Chromatogr. A 2000, 874, 27–43.
- Giovanna, C.; Carmine, C.; Stefano, C.; Matteo, G. β-N-acetylglucosaminidase and β-galactosidase from aleurone layers of resting wheat grains. Phytochemistry1985, 24(7), 1465–9.
- Akolkar, S.K.; Sajgure, A.; Lele, S.S. Lactase production from Lactobacillus acidophilus. World J. Microbiol. Biotechnol. Oct 2005, 21(6–7), 1119–22.
- Benevides, C.C.P.; Roberto, M.; Lorena, B.; et al. Ion exchange using poorly activated supports: an easy way for purification of large proteins. J. Chromatogr. A 2004, 1034, 155–9.
- Diaz, M.; Pedregosa, A.M.; de Lucas, J.R.; Torralba, S.; Monistrol, I.F.; Laborda, F. Purification and properties
of β-galactosidase from Aspergillus nidulans. Microbiol. SEM1996, 12, 585–92. - Gargova, S.; Pishtijski, I.; Stoilova, I. Purification and properties of β-galactosidase from Aspergillus oryzae. Biotechnol.Eq. 1995, 9, 47–51.
The authors are with the Food and Fermentation Technology Department, Institute of Chemical Technology, University of Mumbai, Nathalal Parikh Marg, Matunga, Mumbai 400 019, India; tel.: +95 22 24145616; fax: +95 22 24145614; e-mail: [email protected]. The authors would like to thank the University Grants Commission for providing financial assistance.